CD103 and CD39 coexpression identifies neoantigen-specific cytotoxic T cells in colorectal cancers with low mutation burden
- PMID: 36792124
- PMCID: PMC9933759
- DOI: 10.1136/jitc-2022-005887
CD103 and CD39 coexpression identifies neoantigen-specific cytotoxic T cells in colorectal cancers with low mutation burden
Abstract
Background: Expression of CD103 and CD39 has been found to pinpoint tumor-reactive CD8+ T cells in a variety of solid cancers. We aimed to investigate whether these markers specifically identify neoantigen-specific T cells in colorectal cancers (CRCs) with low mutation burden.
Experimental design: Whole-exome and RNA sequencing of 11 mismatch repair-proficient (MMR-proficient) CRCs and corresponding healthy tissues were performed to determine the presence of putative neoantigens. In parallel, tumor-infiltrating lymphocytes (TILs) were cultured from the tumor fragments and, in parallel, CD8+ T cells were flow-sorted from their respective tumor digests based on single or combined expression of CD103 and CD39. Each subset was expanded and subsequently interrogated for neoantigen-directed reactivity with synthetic peptides. Neoantigen-directed reactivity was determined by flow cytometric analyses of T cell activation markers and ELISA-based detection of IFN-γ and granzyme B release. Additionally, imaging mass cytometry was applied to investigate the localization of CD103+CD39+ cytotoxic T cells in tumors.
Results: Neoantigen-directed reactivity was only encountered in bulk TIL populations and CD103+CD39+ (double positive, DP) CD8+ T cell subsets but never in double-negative or single-positive subsets. Neoantigen-reactivity detected in bulk TIL but not in DP CD8+ T cells could be attributed to CD4+ T cells. CD8+ T cells that were located in direct contact with cancer cells in tumor tissues were enriched for CD103 and CD39 expression.
Conclusion: Coexpression of CD103 and CD39 is characteristic of neoantigen-specific CD8+ T cells in MMR-proficient CRCs with low mutation burden. The exploitation of these subsets in the context of adoptive T cell transfer or engineered T cell receptor therapies is a promising avenue to extend the benefits of immunotherapy to an increasing number of CRC patients.
Keywords: Antigens; CD8-Positive T-Lymphocytes; Gastrointestinal Neoplasms; Immunotherapy; Lymphocytes, Tumor-Infiltrating.
© Author(s) (or their employer(s)) 2023. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ.
Conflict of interest statement
Competing interests: TD and RD disclose that they submitted a patent regarding therapeutic and diagnostic use of the CD103+CD39+ CD8+ T cells in cancer patients. The other authors declare they have no competing interests.
Figures

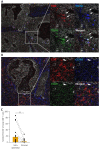
References
-
- Overman MJ, McDermott R, Leach JL, et al. Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (checkmate 142): an open-label, multicentre, phase 2 study. Lancet Oncol 2017;18:1182–91. 10.1016/S1470-2045(17)30422-9 - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
