Digestive enzymes and sphingomyelinase D in spiders without venom (Uloboridae)
- PMID: 36792649
- PMCID: PMC9932164
- DOI: 10.1038/s41598-023-29828-x
Digestive enzymes and sphingomyelinase D in spiders without venom (Uloboridae)
Abstract
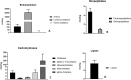
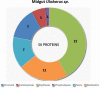
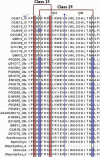
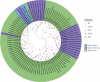
Spiders have distinct predatory behaviours selected along Araneae's evolutionary history but are mainly based on the use of venom for prey paralysis. Uloboridae spiders have lost their venom glands secondarily during evolution. Because of this, they immobilise their prey by extensively wrapping, and digestion starts with the addition of digestive fluid. During the extra-oral digestion, the digestive fluid liquefies both the prey and the AcSp2 spidroins from the web fibres. Despite the efficiency of this process, the cocktail of enzymes involved in digestion in Uloboridae spiders remains unknown. In this study, the protein content in the midgut of Uloborus sp. was evaluated through enzymatic, proteomic, and phylogenetic analysis. Hydrolases such as peptidases (endo and exopeptidases: cysteine, serine, and metallopeptidases), carbohydrases (alpha-amylase, chitinase, and alpha-mannosidase), and lipases were biochemically assayed, and 50 proteins (annotated as enzymes, structural proteins, and toxins) were identified, evidencing the identity between the digestive enzymes present in venomous and non-venomous spiders. Even enzymes thought to be unique to venom, including enzymes such as sphingomyelinase D, were found in the digestive system of non-venomous spiders, suggesting a common origin between digestive enzymes and enzymes present in venoms. This is the first characterization of the molecules involved in the digestive process and the midgut protein content of a non-venomous spider.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Beyond venomous fangs: Uloboridae spiders have lost their venom but not their toxicity.BMC Biol. 2025 Jun 13;23(1):159. doi: 10.1186/s12915-025-02248-1. BMC Biol. 2025. PMID: 40506724 Free PMC article.
-
The deep-rooted origin of disulfide-rich spider venom toxins.Elife. 2023 Feb 9;12:e83761. doi: 10.7554/eLife.83761. Elife. 2023. PMID: 36757362 Free PMC article.
-
Characterisation of protein families in spider digestive fluids and their role in extra-oral digestion.BMC Genomics. 2017 Aug 10;18(1):600. doi: 10.1186/s12864-017-3987-9. BMC Genomics. 2017. PMID: 28797246 Free PMC article.
-
Recent advances in the understanding of brown spider venoms: From the biology of spiders to the molecular mechanisms of toxins.Toxicon. 2014 Jun;83:91-120. doi: 10.1016/j.toxicon.2014.02.023. Epub 2014 Mar 11. Toxicon. 2014. PMID: 24631373 Review.
-
The biology and evolution of spider venoms.Biol Rev Camb Philos Soc. 2022 Feb;97(1):163-178. doi: 10.1111/brv.12793. Epub 2021 Aug 28. Biol Rev Camb Philos Soc. 2022. PMID: 34453398 Review.
Cited by
-
Diversity and Functionality of Bacteria Associated with Different Tissues of Spider Heteropoda venatoria Revealed through Integration of High-Throughput Sequencing and Culturomics Approaches.Microb Ecol. 2024 May 4;87(1):67. doi: 10.1007/s00248-024-02383-2. Microb Ecol. 2024. PMID: 38703220 Free PMC article.
-
A Brief Overview of the Toxic Sphingomyelinase Ds of Brown Recluse Spider Venom and Other Organisms and Simple Methods To Detect Production of Its Signature Cyclic Ceramide Phosphate.Mol Pharmacol. 2024 Feb 15;105(3):144-154. doi: 10.1124/molpharm.123.000709. Mol Pharmacol. 2024. PMID: 37739813 Free PMC article.
-
Enlightening the toxinological dark matter of spider venom enzymes.NPJ Biodivers. 2024 Sep 13;3(1):25. doi: 10.1038/s44185-024-00058-2. NPJ Biodivers. 2024. PMID: 39271930 Free PMC article.
-
Spider venom potency exhibits phylogenetic prey specificity but does not trade-off with body size or silk use in prey capture.Biol Lett. 2025 May;21(5):20250133. doi: 10.1098/rsbl.2025.0133. Epub 2025 May 21. Biol Lett. 2025. PMID: 40393515
-
Structural and Energetic Evidence Supports the Non-Covalent Phosphate Cyclization by the Class II Phospholipase D from Loxosceles intermedia.Toxins (Basel). 2025 Feb 27;17(3):111. doi: 10.3390/toxins17030111. Toxins (Basel). 2025. PMID: 40137884 Free PMC article.
References
-
- Cohen AC. Extra-Oral Digestion in Predaceous Terrestrial Arthropoda. Annu Rev Entomol. 1995;40:85–103. doi: 10.1146/annurev.en.40.010195.000505. - DOI
-
- Foelix, R. Biology of spiders. (OUP USA, 2011).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

