Overdominant expression of genes plays a key role in root growth of tobacco hybrids
- PMID: 36798711
- PMCID: PMC9927235
- DOI: 10.3389/fpls.2023.1107550
Overdominant expression of genes plays a key role in root growth of tobacco hybrids
Abstract
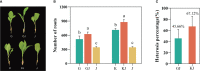
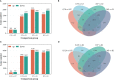
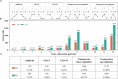
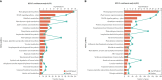
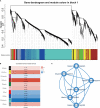
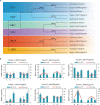
Heterosis has greatly improved the yield and quality of crops. However, previous studies often focused on improving the yield and quality of the shoot system, while research on the root system was neglected. We determined the root numbers of 12 F1 hybrids, all of which showed strong heterosis, indicating that tobacco F1 hybrids have general heterosis. To understand its molecular mechanism, we selected two hybrids with strong heterosis, GJ (G70 × Jiucaiping No.2) and KJ (K326 × Jiucaiping No.2), and their parents for transcriptome analysis. There were 84.22% and 90.25% of the differentially expressed genes were overdominantly expressed. The enrichment analysis of these overdominantly expressed genes showed that "Plant hormone signal transduction", "Phenylpropanoid biosynthesis", "MAPK signaling pathway - plant", and "Starch and sucrose metabolism" pathways were associated with root development. We focused on the analysis of the biosynthetic pathways of auxin(AUX), cytokinins(CTK), abscisic acid(ABA), ethylene(ET), and salicylic acid(SA), suggesting that overdominant expression of these hormone signaling pathway genes may enhance root development in hybrids. In addition, Nitab4.5_0011528g0020、Nitab4.5_0003282g0020、Nitab4.5_0004384g0070 may be the genes involved in root growth. Genome-wide comparative transcriptome analysis enhanced our understanding of the regulatory network of tobacco root development and provided new ideas for studying the molecular mechanisms of tobacco root development.
Keywords: WGCNA; heterosis; overdominant expression; root system; transcriptomics.
Copyright © 2023 Pi, Huang, Luo, Zeng, Mo, Duan and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Non-additive expression genes play a critical role in leaf vein ratio heterosis in Nicotiana tabacum L.BMC Genomics. 2024 Oct 3;25(1):924. doi: 10.1186/s12864-024-10821-1. BMC Genomics. 2024. PMID: 39363277 Free PMC article.
-
Comparative transcriptome analysis of inbred lines and contrasting hybrids reveals overdominance mediate early biomass vigor in hybrid cotton.BMC Genomics. 2020 Feb 10;21(1):140. doi: 10.1186/s12864-020-6561-9. BMC Genomics. 2020. PMID: 32041531 Free PMC article.
-
Overdominance at the Gene Expression Level Plays a Critical Role in the Hybrid Root Growth of Brassica napus.Int J Mol Sci. 2021 Aug 26;22(17):9246. doi: 10.3390/ijms22179246. Int J Mol Sci. 2021. PMID: 34502153 Free PMC article.
-
Comparative transcriptome analysis between inbred lines and hybrids provides molecular insights into K+ content heterosis of tobacco (Nicotiana tabacum L.).Front Plant Sci. 2022 Aug 5;13:940787. doi: 10.3389/fpls.2022.940787. eCollection 2022. Front Plant Sci. 2022. PMID: 35991430 Free PMC article.
-
Transcriptome analysis reveals the key role of overdominant expression of photosynthetic and respiration-related genes in the formation of tobacco(Nicotiana tabacum L.) biomass heterosis.BMC Genomics. 2024 Jun 14;25(1):598. doi: 10.1186/s12864-024-10507-8. BMC Genomics. 2024. PMID: 38877410 Free PMC article.
Cited by
-
Non-additive expression genes play a critical role in leaf vein ratio heterosis in Nicotiana tabacum L.BMC Genomics. 2024 Oct 3;25(1):924. doi: 10.1186/s12864-024-10821-1. BMC Genomics. 2024. PMID: 39363277 Free PMC article.
-
Comparative transcriptome and hormone analyses of roots in apple among three rootstocks with different rooting abilities.PeerJ. 2024 Oct 14;12:e18244. doi: 10.7717/peerj.18244. eCollection 2024. PeerJ. 2024. PMID: 39421411 Free PMC article.
-
Construction and evaluation of a model for efficient identification of photothermal sensitivity of tobacco cultivars based on agronomic traits.Sci Rep. 2024 Nov 13;14(1):27918. doi: 10.1038/s41598-024-78877-3. Sci Rep. 2024. PMID: 39537678 Free PMC article.
References
-
- Belda-Palazon B., Gonzalez-Garcia M. P., Lozano-Juste J., Coego A., Antoni R., Julian J., et al. . (2018). Pyl8 mediates aba perception in the root through non-cell-autonomous and ligand-stabilization-based mechanisms. Proc. Natl. Acad. Sci. U S 115, E11857–E11863. doi: 10.1073/pnas.1815410115 - DOI - PMC - PubMed
-
- Cuadrado A., Cánovas J., Navarrete M. H. (1987). Influence of cell size on differentiation of root meristem cells. Environ. Exp. Bot. 27, 273–277. doi: 10.1016/0098-8472(87)90036-0 - DOI
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

