BRG1 HSA domain interactions with BCL7 proteins are critical for remodeling and gene expression
- PMID: 36801810
- PMCID: PMC9939006
- DOI: 10.26508/lsa.202201770
BRG1 HSA domain interactions with BCL7 proteins are critical for remodeling and gene expression
Abstract
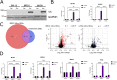
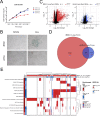
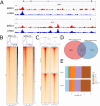
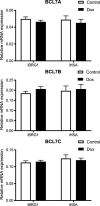
The SWI/SNF complex remodels chromatin in an ATP-dependent manner through the subunits BRG1 and BRM. Chromatin remodeling alters nucleosome structure to change gene expression; however, aberrant remodeling can result in cancer. We identified BCL7 proteins as critical SWI/SNF members that drive BRG1-dependent gene expression changes. BCL7s have been implicated in B-cell lymphoma, but characterization of their functional role within the SWI/SNF complex has been limited. This study implicates their function alongside BRG1 to drive large-scale changes in gene expression. Mechanistically, the BCL7 proteins bind to the HSA domain of BRG1 and require this domain for binding to chromatin. BRG1 proteins without the HSA domain fail to interact with the BCL7 proteins and have severely reduced chromatin remodeling activity. These results link the HSA domain and the formation of a functional SWI/SNF remodeling complex through the interaction with BCL7 proteins. These data highlight the importance of correct formation of the SWI/SNF complex to drive critical biological functions, as losses of individual accessory members or protein domains can cause loss of complex function.
© 2023 Dietrich et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




Similar articles
-
Cdx2 Regulates Gene Expression through Recruitment of Brg1-associated Switch-Sucrose Non-fermentable (SWI-SNF) Chromatin Remodeling Activity.J Biol Chem. 2017 Feb 24;292(8):3389-3399. doi: 10.1074/jbc.M116.752774. Epub 2017 Jan 12. J Biol Chem. 2017. PMID: 28082674 Free PMC article.
-
Novel Interactions between the Human T-Cell Leukemia Virus Type 1 Antisense Protein HBZ and the SWI/SNF Chromatin Remodeling Family: Implications for Viral Life Cycle.J Virol. 2019 Jul 30;93(16):e00412-19. doi: 10.1128/JVI.00412-19. Print 2019 Aug 15. J Virol. 2019. PMID: 31142665 Free PMC article.
-
Glioma tumor suppressor candidate region gene 1 (GLTSCR1) and its paralog GLTSCR1-like form SWI/SNF chromatin remodeling subcomplexes.J Biol Chem. 2018 Mar 16;293(11):3892-3903. doi: 10.1074/jbc.RA117.001065. Epub 2018 Jan 26. J Biol Chem. 2018. PMID: 29374058 Free PMC article.
-
Chromatin remodeling during glucocorticoid receptor regulated transactivation.Biochim Biophys Acta. 2012 Jul;1819(7):716-26. doi: 10.1016/j.bbagrm.2012.02.019. Epub 2012 Mar 6. Biochim Biophys Acta. 2012. PMID: 22425674 Free PMC article. Review.
-
BAFfling pathologies: Alterations of BAF complexes in cancer.Cancer Lett. 2018 Apr 10;419:266-279. doi: 10.1016/j.canlet.2018.01.046. Epub 2018 Jan 31. Cancer Lett. 2018. PMID: 29374542 Review.
Cited by
-
RNA Polymerase II coordinates histone deacetylation at active promoters.bioRxiv [Preprint]. 2024 Sep 17:2024.09.17.613553. doi: 10.1101/2024.09.17.613553. bioRxiv. 2024. Update in: Sci Adv. 2025 Feb 07;11(6):eadt3037. doi: 10.1126/sciadv.adt3037. PMID: 39345547 Free PMC article. Updated. Preprint.
-
Advances in epigenetic therapies for B-cell non-hodgkin lymphoma.Ann Hematol. 2024 Dec;103(12):5085-5101. doi: 10.1007/s00277-024-06131-x. Epub 2024 Dec 9. Ann Hematol. 2024. PMID: 39652169 Review.
-
RNA polymerase II coordinates histone deacetylation at active promoters.Sci Adv. 2025 Feb 7;11(6):eadt3037. doi: 10.1126/sciadv.adt3037. Epub 2025 Feb 5. Sci Adv. 2025. PMID: 39908363 Free PMC article.
-
The role of SMARCA4 in lung cancer.Sci Rep. 2025 Aug 5;15(1):28605. doi: 10.1038/s41598-025-13913-4. Sci Rep. 2025. PMID: 40764807 Free PMC article. Review.
References
-
- Baliñas-Gavira C, Rodríguez MI, Andrades A, Cuadros M, Álvarez-Pérez JC, Álvarez-Prado ÁF, Yébenes VGd, Sánchez-Hernández S, Fernández-Vigo E, Muñoz J, et al. (2020) Frequent mutations in the amino-terminal domain of BCL7A impair its tumor suppressor role in DLBCL. Leukemia 34: 2722–2735. 10.1038/s41375-020-0919-5 - DOI - PubMed
-
- Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: A practical and powerful approach to multiple testing. J R Stat Soc Ser B (Methodological) 57: 289–300. 10.1111/j.2517-6161.1995.tb02031.x - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous
