An open label, non-randomized study assessing a prebiotic fiber intervention in a small cohort of Parkinson's disease participants
- PMID: 36801916
- PMCID: PMC9938693
- DOI: 10.1038/s41467-023-36497-x
An open label, non-randomized study assessing a prebiotic fiber intervention in a small cohort of Parkinson's disease participants
Abstract
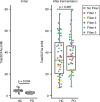
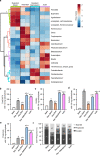
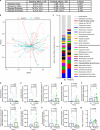
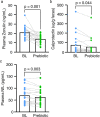
A pro-inflammatory intestinal microbiome is characteristic of Parkinson's disease (PD). Prebiotic fibers change the microbiome and this study sought to understand the utility of prebiotic fibers for use in PD patients. The first experiments demonstrate that fermentation of PD patient stool with prebiotic fibers increased the production of beneficial metabolites (short chain fatty acids, SCFA) and changed the microbiota demonstrating the capacity of PD microbiota to respond favorably to prebiotics. Subsequently, an open-label, non-randomized study was conducted in newly diagnosed, non-medicated (n = 10) and treated PD participants (n = 10) wherein the impact of 10 days of prebiotic intervention was evaluated. Outcomes demonstrate that the prebiotic intervention was well tolerated (primary outcome) and safe (secondary outcome) in PD participants and was associated with beneficial biological changes in the microbiota, SCFA, inflammation, and neurofilament light chain. Exploratory analyses indicate effects on clinically relevant outcomes. This proof-of-concept study offers the scientific rationale for placebo-controlled trials using prebiotic fibers in PD patients. ClinicalTrials.gov Identifier: NCT04512599.
© 2023. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
The prebiotic bar was provided by BetterBiotics Inc which is co-owned by Drs. Keshavarzian, Hamaker, and Sedghi. A provisional patent is pending for the prebiotic mixture used in the bar (Patent Applicant: Purdue Research Foundation, Inventors: Drs. Hamaker, Cantu-Jungles, Keshavarzian, Application Number: 69548-02, Status: Pending, Patent covers the use of prebiotics to improve health). Drs. Keshavarzian, Hamaker, Sedghi, and Cantu-Jungles were not involved in subject recruitment, subject assessments, or statistical analysis of the data. The remaining authors declare no competing interests.
Figures
References
-
- Castillo-Alvarez F, Marzo-Sola ME. Role of the gut microbiota in the development of various neurological diseases. Neurologia. 2019;37:492–498. - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

