Identification of global inhibitors of cellular glycosylation
- PMID: 36804936
- PMCID: PMC9941569
- DOI: 10.1038/s41467-023-36598-7
Identification of global inhibitors of cellular glycosylation
Abstract
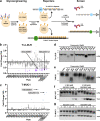
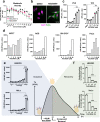
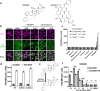
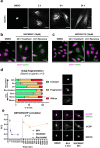
Small molecule inhibitors of glycosylation enzymes are valuable tools for dissecting glycan functions and potential drug candidates. Screening for inhibitors of glycosyltransferases are mainly performed by in vitro enzyme assays with difficulties moving candidates to cells and animals. Here, we circumvent this by employing a cell-based screening assay using glycoengineered cells expressing tailored reporter glycoproteins. We focused on GalNAc-type O-glycosylation and selected the GalNAc-T11 isoenzyme that selectively glycosylates endocytic low-density lipoprotein receptor (LDLR)-related proteins as targets. Our screen of a limited small molecule compound library did not identify selective inhibitors of GalNAc-T11, however, we identify two compounds that broadly inhibited Golgi-localized glycosylation processes. These compounds mediate the reversible fragmentation of the Golgi system without affecting secretion. We demonstrate how these inhibitors can be used to manipulate glycosylation in cells to induce expression of truncated O-glycans and augment binding of cancer-specific Tn-glycoprotein antibodies and to inhibit expression of heparan sulfate and binding and infection of SARS-CoV-2.
© 2023. The Author(s).
Conflict of interest statement
University of Copenhagen has filed a patent application covering the design of the cell-based screening assay for inhibitors described here, which is currently pending. Application numbers are: EP3455635A1, PCT/EP2017/061385, USPTO 20190330601, the inventors are: Eric Bennett, Yoshiki Narimatsu (Y.N.), Catharina Steentoft, Zhang Yang, Ulla Mandel, and Henrik Clausen (H.C.). GlycoDisplay Aps, Copenhagen, Denmark, has obtained a license to the field of the patent application. Y.N. and H.C. are co-founders of GlycoDisplay Aps and hold ownerships in the company as well as served as unpaid consultants. The remaining authors declare no competing interests.
Figures


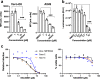
References
-
- Esko, J. D., Bertozzi, C. & Schnaar, R. L. in Essentials of Glycobiology (eds Varki, A. et al.) 701–712 (Cold Spring Harbor Laboratory Press, 2015). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

