Non-canonical pattern recognition of a pathogen-derived metabolite by a nuclear hormone receptor identifies virulent bacteria in C. elegans
- PMID: 36804958
- PMCID: PMC10101930
- DOI: 10.1016/j.immuni.2023.01.027
Non-canonical pattern recognition of a pathogen-derived metabolite by a nuclear hormone receptor identifies virulent bacteria in C. elegans
Abstract
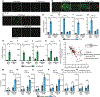
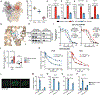
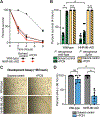
Distinguishing infectious pathogens from harmless microorganisms is essential for animal health. The mechanisms used to identify infectious microbes are not fully understood, particularly in metazoan hosts that eat bacteria as their food source. Here, we characterized a non-canonical pattern-recognition system in Caenorhabditis elegans (C. elegans) that assesses the relative threat of virulent Pseudomonas aeruginosa (P. aeruginosa) to activate innate immunity. We discovered that the innate immune response in C. elegans was triggered by phenazine-1-carboxamide (PCN), a toxic metabolite produced by pathogenic strains of P. aeruginosa. We identified the nuclear hormone receptor NHR-86/HNF4 as the PCN sensor in C. elegans and validated that PCN bound to the ligand-binding domain of NHR-86/HNF4. Activation of NHR-86/HNF4 by PCN directly engaged a transcriptional program in intestinal epithelial cells that protected against P. aeruginosa. Thus, a bacterial metabolite is a pattern of pathogenesis surveilled by nematodes to identify a pathogen in its bacterial diet.
Keywords: Caenorhabditis elegans; Pseudomonas aeruginosa; host-pathogen interactions; innate immunity; intestinal epithelial cells; nuclear hormone receptor; pathogen sensing; pattern recognition receptor; patterns of pathogenesis; phenazines.
Copyright © 2023 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures

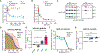

Comment in
-
Pathogen metabolite checkpoint: NHR on guard.Immunity. 2023 Apr 11;56(4):744-746. doi: 10.1016/j.immuni.2023.03.004. Immunity. 2023. PMID: 37044064
Similar articles
-
The nuclear hormone receptor NHR-86 controls anti-pathogen responses in C. elegans.PLoS Genet. 2019 Jan 22;15(1):e1007935. doi: 10.1371/journal.pgen.1007935. eCollection 2019 Jan. PLoS Genet. 2019. PMID: 30668573 Free PMC article.
-
Novel Immune Modulators Enhance Caenorhabditis elegans Resistance to Multiple Pathogens.mSphere. 2021 Jan 6;6(1):e00950-20. doi: 10.1128/mSphere.00950-20. mSphere. 2021. PMID: 33408224 Free PMC article.
-
Pathogen metabolite checkpoint: NHR on guard.Immunity. 2023 Apr 11;56(4):744-746. doi: 10.1016/j.immuni.2023.03.004. Immunity. 2023. PMID: 37044064
-
Modeling Host-Pathogen Interactions in C. elegans: Lessons Learned from Pseudomonas aeruginosa Infection.Int J Mol Sci. 2024 Jun 27;25(13):7034. doi: 10.3390/ijms25137034. Int J Mol Sci. 2024. PMID: 39000143 Free PMC article. Review.
-
Local and long-range activation of innate immunity by infection and damage in C. elegans.Curr Opin Immunol. 2016 Feb;38:1-7. doi: 10.1016/j.coi.2015.09.005. Epub 2015 Oct 28. Curr Opin Immunol. 2016. PMID: 26517153 Review.
Cited by
-
Nuclear hormone receptor NHR-49 is an essential regulator of stress resilience and healthy aging in Caenorhabditis elegans.Front Physiol. 2023 Aug 14;14:1241591. doi: 10.3389/fphys.2023.1241591. eCollection 2023. Front Physiol. 2023. PMID: 37645565 Free PMC article. Review.
-
Mediator subunit MDT-15 promotes expression of propionic acid breakdown genes to prevent embryonic lethality in Caenorhabditis elegans.G3 (Bethesda). 2023 Jun 1;13(6):jkad087. doi: 10.1093/g3journal/jkad087. G3 (Bethesda). 2023. PMID: 37075089 Free PMC article.
-
Antagonism between neuropeptides and monoamines in a distributed circuit for pathogen avoidance.Cell Rep. 2024 Apr 23;43(4):114042. doi: 10.1016/j.celrep.2024.114042. Epub 2024 Apr 3. Cell Rep. 2024. PMID: 38573858 Free PMC article.
-
Patterns of pathogenesis in innate immunity: insights from C. elegans.Nat Rev Immunol. 2025 Apr 17. doi: 10.1038/s41577-025-01167-0. Online ahead of print. Nat Rev Immunol. 2025. PMID: 40247006 Review.
-
Caenorhabditis elegans RIG-I-like receptor DRH-1 signals via CARDs to activate antiviral immunity in intestinal cells.Proc Natl Acad Sci U S A. 2024 Jul 16;121(29):e2402126121. doi: 10.1073/pnas.2402126121. Epub 2024 Jul 9. Proc Natl Acad Sci U S A. 2024. PMID: 38980902 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- P40 OD010440/OD/NIH HHS/United States
- R21 AI149716/AI/NIAID NIH HHS/United States
- R01 AI130289/AI/NIAID NIH HHS/United States
- R01 GM135919/GM/NIGMS NIH HHS/United States
- R21 AI163430/AI/NIAID NIH HHS/United States
- T32 AI095213/AI/NIAID NIH HHS/United States
- R01 AI159159/AI/NIAID NIH HHS/United States
- T32 GM107000/GM/NIGMS NIH HHS/United States
- F30 AI150127/AI/NIAID NIH HHS/United States
- R01 AI150478/AI/NIAID NIH HHS/United States
- F30 DK127690/DK/NIDDK NIH HHS/United States
- T32 AI132152/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

