Identification of RimR2 as a positive pathway-specific regulator of rimocidin biosynthesis in Streptomyces rimosus M527
- PMID: 36810073
- PMCID: PMC9942304
- DOI: 10.1186/s12934-023-02039-9
Identification of RimR2 as a positive pathway-specific regulator of rimocidin biosynthesis in Streptomyces rimosus M527
Abstract
Background: Streoptomyces rimosus M527 is a producer of the polyene macrolide rimocidin which shows activity against various plant pathogenic fungi. Notably, the regulatory mechanisms underlying rimocidin biosynthesis are yet to be elucidated.
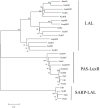
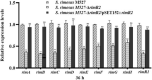
Results: In this study, using domain structure and amino acid alignment and phylogenetic tree construction, rimR2, which located in the rimocidin biosynthetic gene cluster, was first found and identified as a larger ATP-binding regulators of the LuxR family (LAL) subfamily regulator. The rimR2 deletion and complementation assays were conducted to explore its role. Mutant M527-ΔrimR2 lost its ability to produce rimocidin. Complementation of M527-ΔrimR2 restored rimocidin production. The five recombinant strains, M527-ER, M527-KR, M527-21R, M527-57R, and M527-NR, were constructed by overexpressing rimR2 gene using the promoters permE*, kasOp*, SPL21, SPL57, and its native promoter, respectively, to improve rimocidin production. M527-KR, M527-NR, and M527-ER exhibited 81.8%, 68.1%, and 54.5% more rimocidin production, respectively, than the wild-type (WT) strain, while recombinant strains M527-21R and M527-57R exhibited no obvious differences in rimocidin production compared with the WT strain. RT-PCR assays revealed that the transcriptional levels of the rim genes were consistent with the changes in rimocidin production in the recombinant strains. Using electrophoretic mobility shift assays, we confirmed that RimR2 can bind to the promoter regions of rimA and rimC.
Conclusion: A LAL regulator RimR2 was identified as a positive specific-pathway regulator of rimocidin biosynthesis in M527. RimR2 regulates the rimocidin biosynthesis by influencing the transcriptional levels of rim genes and binding to the promoter regions of rimA and rimC.
Keywords: LAL regulator; RimR2; Rimocidin; Streptomyces rimosus.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





Similar articles
-
Transposon-based identification of genes involved in the rimocidin biosynthesis in Streptomyces rimosus M527.World J Microbiol Biotechnol. 2023 Oct 28;39(12):359. doi: 10.1007/s11274-023-03814-x. World J Microbiol Biotechnol. 2023. PMID: 37891332
-
RS24090, a TetR family transcriptional repressor, negatively affects the rimocidin biosynthesis in Streptomyces rimosus M527.Int J Biol Macromol. 2025 Jan;285:138043. doi: 10.1016/j.ijbiomac.2024.138043. Epub 2024 Nov 26. Int J Biol Macromol. 2025. PMID: 39592035
-
Sequential improvement of rimocidin production in Streptomyces rimosus M527 by introduction of cumulative drug-resistance mutations.J Ind Microbiol Biotechnol. 2019 May;46(5):697-708. doi: 10.1007/s10295-019-02146-w. Epub 2019 Jan 29. J Ind Microbiol Biotechnol. 2019. PMID: 30697650
-
Identification of a gene from Streptomyces rimosus M527 negatively affecting rimocidin biosynthesis and morphological differentiation.Appl Microbiol Biotechnol. 2020 Dec;104(23):10191-10202. doi: 10.1007/s00253-020-10955-8. Epub 2020 Oct 15. Appl Microbiol Biotechnol. 2020. PMID: 33057790
-
Effects of addition of elicitors on rimocidin biosynthesis in Streptomyces rimosus M527.Appl Microbiol Biotechnol. 2020 May;104(10):4445-4455. doi: 10.1007/s00253-020-10565-4. Epub 2020 Mar 27. Appl Microbiol Biotechnol. 2020. PMID: 32221690
Cited by
-
Identification of a critical gene involved in the biosynthesis of the polyene macrolide lavencidin in Streptomyces lavendulae FRI-5 using the Target-AID (activation-induced cytidine deaminase) base editing technology.Appl Environ Microbiol. 2025 May 21;91(5):e0097524. doi: 10.1128/aem.00975-24. Epub 2025 Apr 22. Appl Environ Microbiol. 2025. PMID: 40261024 Free PMC article.
-
Transposon-based identification of genes involved in the rimocidin biosynthesis in Streptomyces rimosus M527.World J Microbiol Biotechnol. 2023 Oct 28;39(12):359. doi: 10.1007/s11274-023-03814-x. World J Microbiol Biotechnol. 2023. PMID: 37891332
-
Positive regulation of a LuxR family protein, MilO, in mildiomycin biosynthesis.Appl Environ Microbiol. 2025 Jan 31;91(1):e0165424. doi: 10.1128/aem.01654-24. Epub 2024 Dec 23. Appl Environ Microbiol. 2025. PMID: 39714196 Free PMC article.
-
Chalkophomycin Biosynthesis Revealing Unique Enzyme Architecture for a Hybrid Nonribosomal Peptide Synthetase and Polyketide Synthase.Molecules. 2024 Apr 25;29(9):1982. doi: 10.3390/molecules29091982. Molecules. 2024. PMID: 38731473 Free PMC article.
-
Study on the regulatory mechanism of NsdAsr on rimocidin biosynthesis in Streptomyces rimosus M527.Microb Cell Fact. 2025 Jul 10;24(1):162. doi: 10.1186/s12934-025-02784-z. Microb Cell Fact. 2025. PMID: 40640785 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

