Metagenomic and single-cell RNA-Seq survey of the Helicobacter pylori-infected stomach in asymptomatic individuals
- PMID: 36810249
- PMCID: PMC9977493
- DOI: 10.1172/jci.insight.161042
Metagenomic and single-cell RNA-Seq survey of the Helicobacter pylori-infected stomach in asymptomatic individuals
Abstract
Helicobacter pylori colonization of the gastric niche can persist for years in asymptomatic individuals. To deeply characterize the host-microbiota environment in H. pylori-infected (HPI) stomachs, we collected human gastric tissues and performed metagenomic sequencing, single-cell RNA-Seq (scRNA-Seq), flow cytometry, and fluorescent microscopy. HPI asymptomatic individuals had dramatic changes in the composition of gastric microbiome and immune cells compared with noninfected individuals. Metagenomic analysis uncovered pathway alterations related to metabolism and immune response. scRNA-Seq and flow cytometry data revealed that, in contrast to murine stomachs, ILC2s are virtually absent in the human gastric mucosa, whereas ILC3s are the dominant population. Specifically, proportion of NKp44+ ILC3s out of total ILCs were highly increased in the gastric mucosa of asymptomatic HPI individuals, and correlated with the abundance of selected microbial taxa. In addition, CD11c+ myeloid cells and activated CD4+ T cells and B cells were expanded in HPI individuals. B cells of HPI individuals acquired an activated phenotype and progressed into a highly proliferating germinal-center stage and plasmablast maturation, which correlated with the presence of tertiary lymphoid structures within the gastric lamina propria. Our study provides a comprehensive atlas of the gastric mucosa-associated microbiome and immune cell landscape when comparing asymptomatic HPI and uninfected individuals.
Keywords: Bacterial infections; Cellular immune response; Immunology; Microbiology.
Figures

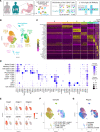





References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials

