Identifying the Best Approximating Model in Bayesian Phylogenetics: Bayes Factors, Cross-Validation or wAIC?
- PMID: 36810802
- PMCID: PMC10276628
- DOI: 10.1093/sysbio/syad004
Identifying the Best Approximating Model in Bayesian Phylogenetics: Bayes Factors, Cross-Validation or wAIC?
Abstract
There is still no consensus as to how to select models in Bayesian phylogenetics, and more generally in applied Bayesian statistics. Bayes factors are often presented as the method of choice, yet other approaches have been proposed, such as cross-validation or information criteria. Each of these paradigms raises specific computational challenges, but they also differ in their statistical meaning, being motivated by different objectives: either testing hypotheses or finding the best-approximating model. These alternative goals entail different compromises, and as a result, Bayes factors, cross-validation, and information criteria may be valid for addressing different questions. Here, the question of Bayesian model selection is revisited, with a focus on the problem of finding the best-approximating model. Several model selection approaches were re-implemented, numerically assessed and compared: Bayes factors, cross-validation (CV), in its different forms (k-fold or leave-one-out), and the widely applicable information criterion (wAIC), which is asymptotically equivalent to leave-one-out cross-validation (LOO-CV). Using a combination of analytical results and empirical and simulation analyses, it is shown that Bayes factors are unduly conservative. In contrast, CV represents a more adequate formalism for selecting the model returning the best approximation of the data-generating process and the most accurate estimates of the parameters of interest. Among alternative CV schemes, LOO-CV and its asymptotic equivalent represented by the wAIC, stand out as the best choices, conceptually and computationally, given that both can be simultaneously computed based on standard Markov chain Monte Carlo runs under the posterior distribution. [Bayes factor; cross-validation; marginal likelihood; model comparison; wAIC.].
© The Author(s) 2023. Published by Oxford University Press on behalf of the Society of Systematic Biologists.
Conflict of interest statement
The author declares no competing interest.
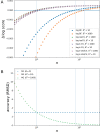
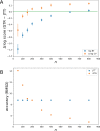
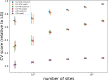
Figures

Similar articles
-
Species delimitation using Bayes factors: simulations and application to the Sceloporus scalaris species group (Squamata: Phrynosomatidae).Syst Biol. 2014 Mar;63(2):119-33. doi: 10.1093/sysbio/syt069. Epub 2013 Nov 20. Syst Biol. 2014. PMID: 24262383
-
Approximating model probabilities in Bayesian information criterion and decision-theoretic approaches to model selection in phylogenetics.Mol Biol Evol. 2011 Jan;28(1):343-9. doi: 10.1093/molbev/msq195. Epub 2010 Jul 29. Mol Biol Evol. 2011. PMID: 20671039 Free PMC article.
-
Bayesian Comparison of Latent Variable Models: Conditional Versus Marginal Likelihoods.Psychometrika. 2019 Sep;84(3):802-829. doi: 10.1007/s11336-019-09679-0. Epub 2019 Jul 11. Psychometrika. 2019. PMID: 31297664
-
Cross-validation to select Bayesian hierarchical models in phylogenetics.BMC Evol Biol. 2016 May 26;16(1):115. doi: 10.1186/s12862-016-0688-y. BMC Evol Biol. 2016. PMID: 27230264 Free PMC article.
-
Searching for convergence in phylogenetic Markov chain Monte Carlo.Syst Biol. 2006 Aug;55(4):553-65. doi: 10.1080/10635150600812544. Syst Biol. 2006. PMID: 16857650
Cited by
-
Measuring the relative contribution to predictive power of modern nucleotide substitution modeling approaches.Bioinform Adv. 2023 Jul 14;3(1):vbad091. doi: 10.1093/bioadv/vbad091. eCollection 2023. Bioinform Adv. 2023. PMID: 37502274 Free PMC article.
-
Annotated Bioinformatic Pipelines for Phylogenomic Placement of Mitochondrial Genomes.Bio Protoc. 2025 Mar 5;15(5):e5232. doi: 10.21769/BioProtoc.5232. eCollection 2025 Mar 5. Bio Protoc. 2025. PMID: 40084070 Free PMC article.
-
Ant backbone phylogeny resolved by modelling compositional heterogeneity among sites in genomic data.Commun Biol. 2024 Jan 17;7(1):106. doi: 10.1038/s42003-024-05793-7. Commun Biol. 2024. PMID: 38233456 Free PMC article.
-
Blouch: Bayesian Linear Ornstein-Uhlenbeck Models for Comparative Hypotheses.Syst Biol. 2024 Nov 29;73(6):1038-1050. doi: 10.1093/sysbio/syae044. Syst Biol. 2024. PMID: 39046734 Free PMC article.
-
Bayesian StairwayPlot for Inferring Single Population Demographic Histories From Site Frequency Spectra.Mol Ecol Resour. 2025 Aug;25(6):e14087. doi: 10.1111/1755-0998.14087. Epub 2025 Feb 26. Mol Ecol Resour. 2025. PMID: 40012349 Free PMC article.
References
-
- Aho K., Derryberry D., Peterson T.. 2014. Model selection for ecologists: the worldviews of AIC and BIC. Ecology 95(3):631–636. - PubMed
-
- Akaike H. 1974. A new look at the statistical model identification. IEEE Trans. Automat. Contr. 19(6):716–723.
-
- Baele G., Lemey P.. 2013. Bayesian evolutionary model testing in the phylogenomics era: matching model complexity with computational efficiency. Bioinformatics 29(16):1970–1979. - PubMed

