Optical photothermal infrared spectroscopy: A novel solution for rapid identification of antimicrobial resistance at the single-cell level via deuterium isotope labeling
- PMID: 36819022
- PMCID: PMC9929359
- DOI: 10.3389/fmicb.2023.1077106
Optical photothermal infrared spectroscopy: A novel solution for rapid identification of antimicrobial resistance at the single-cell level via deuterium isotope labeling
Abstract
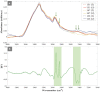
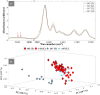
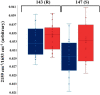
The rise and extensive spread of antimicrobial resistance (AMR) has become a growing concern, and a threat to the environment and human health globally. The majority of current AMR identification methods used in clinical setting are based on traditional microbiology culture-dependent techniques which are time-consuming or expensive to be implemented, thus appropriate antibiotic stewardship is provided retrospectively which means the first line of treatment is to hope that a broad-spectrum antibiotic works. Hence, culture-independent and single-cell technologies are needed to allow for rapid detection and identification of antimicrobial-resistant bacteria and to support a more targeted and effective antibiotic therapy preventing further development and spread of AMR. In this study, for the first time, a non-destructive phenotyping method of optical photothermal infrared (O-PTIR) spectroscopy, coupled with deuterium isotope probing (DIP) and multivariate statistical analysis was employed as a metabolic fingerprinting approach to detect AMR in Uropathogenic Escherichia coli (UPEC) at both single-cell and population levels. Principal component-discriminant function analysis (PC-DFA) of FT-IR and O-PTIR spectral data showed clear clustering patterns as a result of distinctive spectral shifts (C-D signature peaks) originating from deuterium incorporation into bacterial cells, allowing for rapid detection and classification of sensitive and resistant isolates at the single-cell level. Furthermore, the single-frequency images obtained using the C-D signature peak at 2,163 cm-1 clearly displayed the reduced ability of the trimethoprim-sensitive strain for incorporating deuterium when exposed to this antibiotic, compared to the untreated condition. Hence, the results of this study indicated that O-PTIR can be employed as an efficient tool for the rapid detection of AMR at the single-cell level.
Keywords: Uropathogenic Escherichia coli; antimicrobial resistance; infrared spectroscopy; microbiology; single-cell; stable-isotope probing.
Copyright © 2023 Shams, Lima, Xu, Ahmed, Goodacre and Muhamadali.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Application of infrared spectroscopy to study carbon-deuterium kinetics and isotopic spectral shifts at the single-cell level.Spectrochim Acta A Mol Biomol Spectrosc. 2025 Feb 15;327:125374. doi: 10.1016/j.saa.2024.125374. Epub 2024 Nov 3. Spectrochim Acta A Mol Biomol Spectrosc. 2025. PMID: 39522229
-
Raman-Deuterium Isotope Probing for in-situ identification of antimicrobial resistant bacteria in Thames River.Sci Rep. 2017 Nov 30;7(1):16648. doi: 10.1038/s41598-017-16898-x. Sci Rep. 2017. PMID: 29192181 Free PMC article.
-
Campylobacter jejuni Antimicrobial Resistance Profiles and Mechanisms Determined Using a Raman Spectroscopy-Based Metabolomic Approach.Appl Environ Microbiol. 2021 May 26;87(12):e0038821. doi: 10.1128/AEM.00388-21. Epub 2021 May 26. Appl Environ Microbiol. 2021. PMID: 33837016 Free PMC article.
-
Antimicrobial Susceptibility Testing: A Comprehensive Review of Currently Used Methods.Antibiotics (Basel). 2022 Mar 23;11(4):427. doi: 10.3390/antibiotics11040427. Antibiotics (Basel). 2022. PMID: 35453179 Free PMC article. Review.
-
Antimicrobial resistance and ESBL production in uropathogenic Escherichia coli: a systematic review and meta-analysis in Ethiopia.JAC Antimicrob Resist. 2024 May 6;6(3):dlae068. doi: 10.1093/jacamr/dlae068. eCollection 2024 Jun. JAC Antimicrob Resist. 2024. PMID: 38716402 Free PMC article. Review.
Cited by
-
Optical photothermal infrared imaging using metabolic probes in biological systems.bioRxiv [Preprint]. 2025 Jan 15:2024.09.19.613881. doi: 10.1101/2024.09.19.613881. bioRxiv. 2025. Update in: Anal Chem. 2025 Apr 22;97(15):8202-8212. doi: 10.1021/acs.analchem.4c03752. PMID: 39345466 Free PMC article. Updated. Preprint.
-
An optical photothermal infrared investigation of lymph nodal metastases of oral squamous cell carcinoma.Sci Rep. 2024 Jul 11;14(1):16050. doi: 10.1038/s41598-024-66977-z. Sci Rep. 2024. PMID: 38992088 Free PMC article.
-
Bacterial discrimination by Fourier transform infrared spectroscopy, MALDI-mass spectrometry and whole-genome sequencing.Future Microbiol. 2024 Jun 12;19(9):795-810. doi: 10.2217/fmb-2024-0043. Epub 2024 May 29. Future Microbiol. 2024. PMID: 38652264 Free PMC article.
-
Monitoring Phenotype Heterogeneity at the Single-Cell Level within Bacillus Populations Producing Poly-3-hydroxybutyrate by Label-Free Super-resolution Infrared Imaging.Anal Chem. 2023 Dec 5;95(48):17733-17740. doi: 10.1021/acs.analchem.3c03595. Epub 2023 Nov 23. Anal Chem. 2023. PMID: 37997371 Free PMC article.
-
Rapid detection of antimicrobial resistance in methicillin-resistant Staphylococcus aureus using MALDI-TOF mass spectrometry.Front Cell Infect Microbiol. 2023 Nov 23;13:1281155. doi: 10.3389/fcimb.2023.1281155. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 38076465 Free PMC article.
References
-
- Abdi H., Williams L. J. (2010). Principal component analysis. Wiley Interdisciplin. Rev. 2, 433–459. doi: 10.1002/wics.101 - DOI
-
- Afseth N. K., Kohler A. (2012). Extended multiplicative signal correction in vibrational spectroscopy, a tutorial. Chemom. Intell. Lab. Syst. 117, 92–99. doi: 10.1016/j.chemolab.2012.03.004 - DOI
-
- Alsberg B. K., Wade W. G., Goodacre R. (1998). Chemometric analysis of diffuse reflectance-absorbance Fourier transform infrared spectra using rule induction methods: application to the classification of Eubacterium species. Appl. Spectrosc. 52, 823–832. doi: 10.1366/0003702981944562 - DOI
LinkOut - more resources
Full Text Sources

