Induction of primordial germ cell-like cells from common marmoset embryonic stem cells by inhibition of WNT and retinoic acid signaling
- PMID: 36823310
- PMCID: PMC9950483
- DOI: 10.1038/s41598-023-29850-z
Induction of primordial germ cell-like cells from common marmoset embryonic stem cells by inhibition of WNT and retinoic acid signaling
Abstract
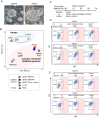
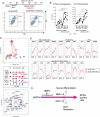
Reconstitution of the germ cell lineage using pluripotent stem cells provides a unique platform to deepen our understanding of the mechanisms underlying germ cell development and to produce functional gametes for reproduction. This study aimed to establish a culture system that induces a robust number of primordial germ cell-like cells (PGCLCs) from common marmoset (Callithrix jacchus) embryonic stem cells. The robust induction was achieved by not only activation of the conserved PGC-inducing signals, WNT and BMP4, but also temporal inhibitions of WNT and retinoic acid signals, which prevent mesodermal and neural differentiation, respectively, during PGCLC differentiation. Many of the gene expression and differentiation properties of common marmoset PGCLCs were similar to those of human PGCLCs, making this culture system a reliable and useful primate model. Finally, we identified PDPN and KIT as surface marker proteins by which PGCLCs can be isolated from embryonic stem cells without genetic manipulation. This study will expand the opportunities for research on germ cell development and production of functional gametes to the common marmoset.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

