PTEN phosphatase inhibits metastasis by negatively regulating the Entpd5/IGF1R pathway through ATF6
- PMID: 36824269
- PMCID: PMC9942123
- DOI: 10.1016/j.isci.2023.106070
PTEN phosphatase inhibits metastasis by negatively regulating the Entpd5/IGF1R pathway through ATF6
Abstract
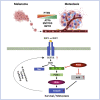
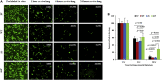
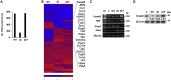
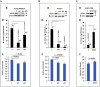
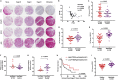
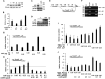
PTEN encodes a tumor suppressor with lipid and protein phosphatase activities whose dysfunction has been implicated in melanomagenesis; less is known about how its phosphatases regulate melanoma metastasis. We demonstrate that PTEN expression negatively correlates with metastatic progression in human melanoma samples and a PTEN-deficient mouse melanoma model. Wildtype PTEN expression inhibited melanoma cell invasiveness and metastasis in a dose-dependent manner, behaviors that specifically required PTEN protein phosphatase activity. PTEN phosphatase activity regulated metastasis through Entpd5. Entpd5 knockdown reduced metastasis and IGF1R levels while promoting ER stress. In contrast, Entpd5 overexpression promoted metastasis and enhanced IGF1R levels while reducing ER stress. Moreover, Entpd5 expression was regulated by the ER stress sensor ATF6. Altogether, our data indicate that PTEN phosphatase activity inhibits metastasis by negatively regulating the Entpd5/IGF1R pathway through ATF6, thereby identifying novel candidate therapeutic targets for the treatment of PTEN mutant melanoma.
Keywords: Cancer; Cell biology; Functional aspects of cell biology.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



Similar articles
-
The ER UDPase ENTPD5 promotes protein N-glycosylation, the Warburg effect, and proliferation in the PTEN pathway.Cell. 2010 Nov 24;143(5):711-24. doi: 10.1016/j.cell.2010.10.010. Epub 2010 Nov 11. Cell. 2010. PMID: 21074248
-
Knockdown of ENTPD5 inhibits tumor metastasis and growth via regulating the GRP78/p-eIF-2α/CHOP pathway in serous ovarian cancer.J Ovarian Res. 2022 Jun 7;15(1):69. doi: 10.1186/s13048-022-00996-0. J Ovarian Res. 2022. PMID: 35668504 Free PMC article.
-
The UDPase ENTPD5 regulates ER stress-associated renal injury by mediating protein N-glycosylation.Cell Death Dis. 2023 Feb 27;14(2):166. doi: 10.1038/s41419-023-05685-4. Cell Death Dis. 2023. PMID: 36849424 Free PMC article.
-
PTEN and Other PtdIns(3,4,5)P3 Lipid Phosphatases in Breast Cancer.Int J Mol Sci. 2020 Dec 2;21(23):9189. doi: 10.3390/ijms21239189. Int J Mol Sci. 2020. PMID: 33276499 Free PMC article. Review.
-
PTEN signaling pathways in melanoma.Oncogene. 2003 May 19;22(20):3113-22. doi: 10.1038/sj.onc.1206451. Oncogene. 2003. PMID: 12789288 Review.
Cited by
-
Development of Personalized Strategies for Precisely Battling Malignant Melanoma.Int J Mol Sci. 2024 May 4;25(9):5023. doi: 10.3390/ijms25095023. Int J Mol Sci. 2024. PMID: 38732242 Free PMC article. Review.
-
PTEN Lipid Phosphatase Activity Suppresses Melanoma Formation by Opposing an AKT/mTOR/FRA1 Signaling Axis.Cancer Res. 2024 Feb 1;84(3):388-404. doi: 10.1158/0008-5472.CAN-23-1730. Cancer Res. 2024. PMID: 38193852 Free PMC article.
-
Reciprocal negative feedback regulation of ATF6α and PTEN promotes prostate cancer progression.Cell Mol Life Sci. 2023 Sep 16;80(10):292. doi: 10.1007/s00018-023-04940-3. Cell Mol Life Sci. 2023. PMID: 37715829 Free PMC article.
-
ATF6α inhibits ΔNp63α expression to promote breast cancer metastasis by the GRP78-AKT1-FOXO3a signaling.Cell Death Dis. 2025 Apr 13;16(1):289. doi: 10.1038/s41419-025-07619-8. Cell Death Dis. 2025. PMID: 40223122 Free PMC article.
-
Arid4a Suppresses Breast Tumor Metastasis by Enhancing MTSS1 Expression via mRNA Stability.Cancer Med. 2025 Mar;14(5):e70732. doi: 10.1002/cam4.70732. Cancer Med. 2025. PMID: 40066676 Free PMC article.
References
-
- Steck P.A., Pershouse M.A., Jasser S.A., Yung W.K., Lin H., Ligon A.H., Langford L.A., Baumgard M.L., Hattier T., Davis T., et al. Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat. Genet. 1997;15:356–362. - PubMed
-
- Li J., Yen C., Liaw D., Podsypanina K., Bose S., Wang S.I., Puc J., Miliaresis C., Rodgers L., McCombie R., et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science. 1997;275:1943–1947. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

