Comparative Modeling and Analysis of Extremophilic D-Ala-D-Ala Carboxypeptidases
- PMID: 36830697
- PMCID: PMC9953012
- DOI: 10.3390/biom13020328
Comparative Modeling and Analysis of Extremophilic D-Ala-D-Ala Carboxypeptidases
Abstract
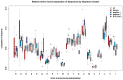
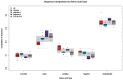
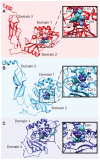
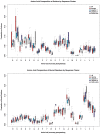
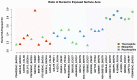
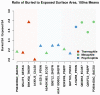
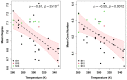
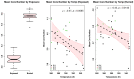
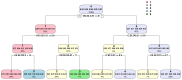
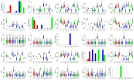
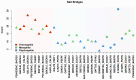
Understanding the molecular adaptations of organisms to extreme environments requires a comparative analysis of protein structure, function, and dynamics across species found in different environmental conditions. Computational studies can be particularly useful in this pursuit, allowing exploratory studies of large numbers of proteins under different thermal and chemical conditions that would be infeasible to carry out experimentally. Here, we perform such a study of the MEROPS family S11, S12, and S13 proteases from psychophilic, mesophilic, and thermophilic bacteria. Using a combination of protein structure prediction, atomistic molecular dynamics, and trajectory analysis, we examine both conserved features and trends across thermal groups. Our findings suggest a number of hypotheses for experimental investigation.
Keywords: MEROPS S11; comparative modeling; extremophiles; machine learning; molecular dynamics; serine protease.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Lonergan Z.R., Nairn B.L., Wang J., Hsu Y.P., Hesse L.E., Beavers W.N., Chazin W.J., Trinidad J.C., Nieuwenhze M.S.V., Giedroc D.P., et al. An Acinetobacter baumannii, zinc-regulated peptidase 650 maintains cell wall integrity during immune-mediated nutrient sequestration. Cell Rep. 2019;26:2009–2018.e6. doi: 10.1016/j.celrep.2019.01.089. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

