Combining Breast Cancer Risk Prediction Models
- PMID: 36831433
- PMCID: PMC9953824
- DOI: 10.3390/cancers15041090
Combining Breast Cancer Risk Prediction Models
Abstract
Accurate risk stratification is key to reducing cancer morbidity through targeted screening and preventative interventions. Multiple breast cancer risk prediction models are used in clinical practice, and often provide a range of different predictions for the same patient. Integrating information from different models may improve the accuracy of predictions, which would be valuable for both clinicians and patients. BRCAPRO is a widely used model that predicts breast cancer risk based on detailed family history information. A major limitation of this model is that it does not consider non-genetic risk factors. To address this limitation, we expand BRCAPRO by combining it with another popular existing model, BCRAT (i.e., Gail), which uses a largely complementary set of risk factors, most of them non-genetic. We consider two approaches for combining BRCAPRO and BCRAT: (1) modifying the penetrance (age-specific probability of developing cancer given genotype) functions in BRCAPRO using relative hazard estimates from BCRAT, and (2) training an ensemble model that takes BRCAPRO and BCRAT predictions as input. Using both simulated data and data from Newton-Wellesley Hospital and the Cancer Genetics Network, we show that the combination models are able to achieve performance gains over both BRCAPRO and BCRAT. In the Cancer Genetics Network cohort, we show that the proposed BRCAPRO + BCRAT penetrance modification model performs comparably to IBIS, an existing model that combines detailed family history with non-genetic risk factors.
Keywords: BCRAT; BRCAPRO; ensemble learning; model aggregation; stacking.
Conflict of interest statement
Giovanni Parmigiani is a member of the Scientific Advisory Board of Ambry Genetics, a commercial provider of cancer susceptibility tests. Commercial use of the BRCAPRO model is licensed by the Dana Farber Cancer Institute as part of the BayesMendel software package. Licensing revenues are used for model and software upgrades. None of the authors derives personal income from BRCAPRO licensing revenues. Kevin Hughes receives Honoraria from Hologic (Surgical implant for radiation planning with breast conservation and wire free breast biopsy) and Myriad Genetics. He has a financial interest in CRA Health (Formerly Hughes RiskApps), which was recently sold to Volpara. CRA Health develops risk assessment models/software with a particular focus on breast cancer and colorectal cancer. Kevin Hughes is a founder of the company. He is the Co-Creator of Ask2Me.Org which is freely available for clinical use and is licensed for commercial use by the Dana Farber Cancer Institute, Harvard University, and the Massachusetts General Hospital (MGH). His interests in CRA Health and Ask2Me.Org were reviewed and are managed by MGH and Partners Health Care in accordance with their conflict of interest policies.
Figures
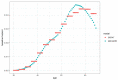

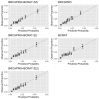
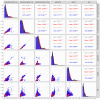
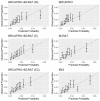
Similar articles
-
10-year performance of four models of breast cancer risk: a validation study.Lancet Oncol. 2019 Apr;20(4):504-517. doi: 10.1016/S1470-2045(18)30902-1. Epub 2019 Feb 21. Lancet Oncol. 2019. PMID: 30799262
-
Breast cancer risk assessment across the risk continuum: genetic and nongenetic risk factors contributing to differential model performance.Breast Cancer Res. 2012 Nov 5;14(6):R144. doi: 10.1186/bcr3352. Breast Cancer Res. 2012. PMID: 23127309 Free PMC article.
-
Variant-specific Mendelian Risk Prediction Model.bioRxiv [Preprint]. 2023 Mar 8:2023.03.06.531363. doi: 10.1101/2023.03.06.531363. bioRxiv. 2023. PMID: 36945459 Free PMC article. Preprint.
-
Recent Enhancements to the Genetic Risk Prediction Model BRCAPRO.Cancer Inform. 2015 May 10;14(Suppl 2):147-57. doi: 10.4137/CIN.S17292. eCollection 2015. Cancer Inform. 2015. PMID: 25983549 Free PMC article. Review.
-
Assessment of the risk of developing breast cancer using the Gail model in Asian females: A systematic review.Heliyon. 2020 Apr 22;6(4):e03794. doi: 10.1016/j.heliyon.2020.e03794. eCollection 2020 Apr. Heliyon. 2020. PMID: 32346636 Free PMC article. Review.
Cited by
-
Deep learning of longitudinal mammogram examinations for breast cancer risk prediction.Pattern Recognit. 2022 Dec;132:108919. doi: 10.1016/j.patcog.2022.108919. Epub 2022 Jul 22. Pattern Recognit. 2022. PMID: 37089470 Free PMC article.
-
Validation of Breast Cancer Risk Models by Race/Ethnicity, Family History and Molecular Subtypes.Cancers (Basel). 2021 Dec 23;14(1):45. doi: 10.3390/cancers14010045. Cancers (Basel). 2021. PMID: 35008209 Free PMC article.
-
Association Between Risk Factors and Major Cancers: Explainable Machine Learning Approach.JMIR Cancer. 2025 May 2;11:e62833. doi: 10.2196/62833. JMIR Cancer. 2025. PMID: 40315870 Free PMC article.
-
Critical Risk Assessment, Diagnosis, and Survival Analysis of Breast Cancer.Diagnostics (Basel). 2024 May 8;14(10):984. doi: 10.3390/diagnostics14100984. Diagnostics (Basel). 2024. PMID: 38786282 Free PMC article.
-
Multiplex Digital Spatial Profiling in Breast Cancer Research: State-of-the-Art Technologies and Applications across the Translational Science Spectrum.Cancers (Basel). 2024 Apr 23;16(9):1615. doi: 10.3390/cancers16091615. Cancers (Basel). 2024. PMID: 38730568 Free PMC article. Review.
References
-
- American Cancer Society Facts and Figures 2020. 2020. [(accessed on 3 May 2020)]. Available online: https://www.cancer.org/research/cancer-facts-statistics/all-cancer-facts....
-
- Cintolo-Gonzalez J.A., Braun D., Blackford A.L., Mazzola E., Acar A., Plichta J.K., Griffin M., Hughes K.S. Breast cancer risk models: A comprehensive overview of existing models, validation, and clinical applications. Breast Cancer Res. Treat. 2017;164:263–284. doi: 10.1007/s10549-017-4247-z. - DOI - PubMed
-
- Gail M.H., Costantino J.P., Pee D., Bondy M., Newman L., Selvan M., Anderson G.L., Malone K.E., Marchbanks P.A., McCaskill-Stevens W., et al. Projecting individualized absolute invasive breast cancer risk in African American women. J. Natl. Cancer Inst. 2007;99:1782–1792. doi: 10.1093/jnci/djm223. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

