In Situ Imaging of O-Linked β-N-Acetylglucosamine Using On-Tissue Hydrolysis and MALDI Mass Spectrometry
- PMID: 36831567
- PMCID: PMC9954453
- DOI: 10.3390/cancers15041224
In Situ Imaging of O-Linked β-N-Acetylglucosamine Using On-Tissue Hydrolysis and MALDI Mass Spectrometry
Abstract
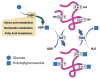
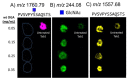
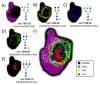
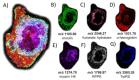
Post-translational O-glycosylation of proteins via the addition of N-acetylglucosamine (O-GlcNAc) is a regulator of many aspects of cellular physiology. Processes driven by perturbed dynamics of O-GlcNAcylation modification have been implicated in cancer development. Variability in O-GlcNAcylation is emerging as a metabolic biomarker of many cancers. Here, we evaluate the use of MALDI-mass spectrometry imaging (MSI) to visualize the location of O-GlcNAcylated proteins in tissue sections by mapping GlcNAc that has been released by the enzymatic hydrolysis of glycoproteins using an O-GlcNAc hydrolase. We use this strategy to monitor O-GlcNAc within hepatic VX2 tumor tissue. We show that increased O-GlcNAc is found within both viable tumor and tumor margin regions, implicating GlcNAc in tumor progression.
Keywords: MALDI; O-GlcNAc; O-glycosylation; biomarkers; in situ digestion; mass spectrometry imaging; ultraviolet photodissociation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
A novel strategy for global mapping of O-GlcNAc proteins and peptides using selective enzymatic deglycosylation, HILIC enrichment and mass spectrometry identification.Talanta. 2017 Jul 1;169:195-202. doi: 10.1016/j.talanta.2017.03.049. Epub 2017 Mar 18. Talanta. 2017. PMID: 28411811
-
Identification of structural and functional O-linked N-acetylglucosamine-bearing proteins in Xenopus laevis oocyte.Mol Cell Proteomics. 2008 Nov;7(11):2229-45. doi: 10.1074/mcp.M700494-MCP200. Epub 2008 Jul 9. Mol Cell Proteomics. 2008. PMID: 18617508
-
O-GlcNAcylation site mapping by (azide-alkyne) click chemistry and mass spectrometry following intensive fractionation of skeletal muscle cells proteins.J Proteomics. 2018 Aug 30;186:83-97. doi: 10.1016/j.jprot.2018.07.005. Epub 2018 Jul 26. J Proteomics. 2018. PMID: 30016717
-
Chemical biology tools to interrogate the roles of O-GlcNAc in immunity.Front Immunol. 2023 Jan 26;13:1089824. doi: 10.3389/fimmu.2022.1089824. eCollection 2022. Front Immunol. 2023. PMID: 36776401 Free PMC article. Review.
-
Chemical arsenal for the study of O-GlcNAc.Molecules. 2011 Feb 28;16(3):1987-2022. doi: 10.3390/molecules16031987. Molecules. 2011. PMID: 21358590 Free PMC article. Review.
Cited by
-
Immunohistochemical Study on O-GlcNAcylation in Diabetic Pathologies: Molecular Mechanisms and Implications.Acta Histochem Cytochem. 2025 Jun 24;58(3):115-121. doi: 10.1267/ahc.25-00022. Epub 2025 Jun 18. Acta Histochem Cytochem. 2025. PMID: 40630795 Free PMC article. Review.
-
Immunohistochemical Study on O-GlcNAcylation in Diabetic Pathologies: Molecular Mechanisms and Implications.Acta Histochem Cytochem. 2025 Jun 24;58(3):115-121. doi: 10.1267/ahc.25-00022. Epub 2025 Jun 18. Acta Histochem Cytochem. 2025. PMID: 40630794 Free PMC article. Review.
References
-
- Herrera H., Dilday T., Uber A., Scott D., Zambrano J.N., Wang M., Angel P.M., Mehta A.S., Drake R.R., Hill E.G., et al. Core-Fucosylated Tetra-Antennary N-Glycan Containing A Single N-Acetyllactosamine Branch Is Associated with Poor Survival Outcome in Breast Cancer. Int. J. Mol. Sci. 2019;20:2528. doi: 10.3390/ijms20102528. - DOI - PMC - PubMed
-
- Briggs M.T., Condina M.R., Ho Y.Y., Everest-Dass A.V., Mittal P., Kaur G., Oehler M.K., Packer N.H., Hoffmann P. MALDI Mass Spectrometry Imaging of Early- and Late-Stage Serous Ovarian Cancer Tissue Reveals Stage-Specific N-Glycans. Proteomics. 2019;19:1800482. doi: 10.1002/pmic.201800482. - DOI - PubMed
-
- Zhang H., Shi X., Liu Y., Wang B., Xu M., Welham N.V., Li L. On-Tissue Amidation of Sialic Acid with Aniline for Sensitive Imaging of Sialylated N-Glycans from FFPE Tissue Sections via MALDI Mass Spectrometry. Anal. Bioanal. Chem. 2022;414:5263–5274. doi: 10.1007/s00216-022-03894-y. - DOI - PMC - PubMed
-
- Boyaval F., Dalebout H., Van Zeijl R., Wang W., Fariña-Sarasqueta A., Lageveen-Kammeijer G.S.M., Boonstra J.J., McDonnell L.A., Wuhrer M., Morreau H., et al. High-Mannose N-Glycans as Malignant Progression Markers in Early-Stage Colorectal Cancer. Cancers. 2022;14:1552. doi: 10.3390/cancers14061552. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

