MicroRNAs and Gene Regulatory Networks Related to Cleft Lip and Palate
- PMID: 36834963
- PMCID: PMC9958963
- DOI: 10.3390/ijms24043552
MicroRNAs and Gene Regulatory Networks Related to Cleft Lip and Palate
Abstract
Cleft lip and palate is one of the most common congenital birth defects and has a complex etiology. Either genetic or environmental factors, or both, are involved at various degrees, and the type and severity of clefts vary. One of the longstanding questions is how environmental factors lead to craniofacial developmental anomalies. Recent studies highlight non-coding RNAs as potential epigenetic regulators in cleft lip and palate. In this review, we will discuss microRNAs, a type of small non-coding RNAs that can simultaneously regulate expression of many downstream target genes, as a causative mechanism of cleft lip and palate in humans and mice.
Keywords: cleft lip; cleft palate; craniofacial development; environmental factor; microRNA.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
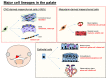

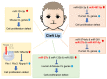

References
-
- Gonseth S., Shaw G., Roy R., Segal M., Asrani K., Rine J., Wiemels J., Marini N. Epigenomic profiling of newborns with isolated orofacial clefts reveals widespread DNA methylation changes and implicates metastable epiallele regions in disease risk. Epigenetics. 2019;14:198–213. doi: 10.1080/15592294.2019.1581591. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

