Genome-Wide Characterization of the PIFs Family in Sweet Potato and Functional Identification of IbPIF3.1 under Drought and Fusarium Wilt Stresses
- PMID: 36835500
- PMCID: PMC9965949
- DOI: 10.3390/ijms24044092
Genome-Wide Characterization of the PIFs Family in Sweet Potato and Functional Identification of IbPIF3.1 under Drought and Fusarium Wilt Stresses
Abstract
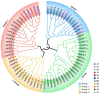
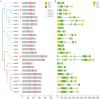
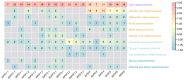
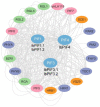
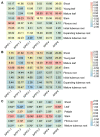
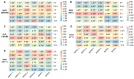
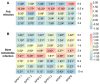
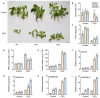
Phytochrome-interacting factors (PIFs) are essential for plant growth, development, and defense responses. However, research on the PIFs in sweet potato has been insufficient to date. In this study, we identified PIF genes in the cultivated hexaploid sweet potato (Ipomoea batatas) and its two wild relatives, Ipomoea triloba, and Ipomoea trifida. Phylogenetic analysis revealed that IbPIFs could be divided into four groups, showing the closest relationship with tomato and potato. Subsequently, the PIFs protein properties, chromosome location, gene structure, and protein interaction network were systematically analyzed. RNA-Seq and qRT-PCR analyses showed that IbPIFs were mainly expressed in stem, as well as had different gene expression patterns in response to various stresses. Among them, the expression of IbPIF3.1 was strongly induced by salt, drought, H2O2, cold, heat, Fusarium oxysporum f. sp. batatas (Fob), and stem nematodes, indicating that IbPIF3.1 might play an important role in response to abiotic and biotic stresses in sweet potato. Further research revealed that overexpression of IbPIF3.1 significantly enhanced drought and Fusarium wilt tolerance in transgenic tobacco plants. This study provides new insights for understanding PIF-mediated stress responses and lays a foundation for future investigation of sweet potato PIFs.
Keywords: Ipomoea batatas; Ipomoea trifida; Ipomoea triloba; PIFs; expression analysis; function analysis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Genome-wide identification, characterisation and expression profile analysis of DEAD-box family genes in sweet potato wild ancestor Ipomoea trifida under abiotic stresses.Genes Genomics. 2020 Mar;42(3):325-335. doi: 10.1007/s13258-019-00910-x. Epub 2020 Jan 1. Genes Genomics. 2020. PMID: 31894476
-
Genome-Wide Identification and Characterization of CDPK Family Reveal Their Involvements in Growth and Development and Abiotic Stress in Sweet Potato and Its Two Diploid Relatives.Int J Mol Sci. 2022 Mar 13;23(6):3088. doi: 10.3390/ijms23063088. Int J Mol Sci. 2022. PMID: 35328509 Free PMC article.
-
Genome-Wide Identification and Expression Analysis of ACTIN Family Genes in the Sweet Potato and Its Two Diploid Relatives.Int J Mol Sci. 2023 Jun 30;24(13):10930. doi: 10.3390/ijms241310930. Int J Mol Sci. 2023. PMID: 37446107 Free PMC article.
-
Current status in whole genome sequencing and analysis of Ipomoea spp.Plant Cell Rep. 2019 Nov;38(11):1365-1371. doi: 10.1007/s00299-019-02464-4. Epub 2019 Aug 29. Plant Cell Rep. 2019. PMID: 31468128 Free PMC article. Review.
-
Multiple biological functions of sporamin related to stress tolerance in sweet potato (Ipomoea batatas Lam).Biotechnol Adv. 2012 Nov-Dec;30(6):1309-17. doi: 10.1016/j.biotechadv.2012.01.022. Epub 2012 Jan 28. Biotechnol Adv. 2012. PMID: 22306516 Review.
Cited by
-
A comprehensive overview of omics-based approaches to enhance biotic and abiotic stress tolerance in sweet potato.Hortic Res. 2024 Jan 12;11(3):uhae014. doi: 10.1093/hr/uhae014. eCollection 2024 Mar. Hortic Res. 2024. PMID: 38464477 Free PMC article.
-
Functions of Plant Phytochrome Signaling Pathways in Adaptation to Diverse Stresses.Int J Mol Sci. 2023 Aug 25;24(17):13201. doi: 10.3390/ijms241713201. Int J Mol Sci. 2023. PMID: 37686008 Free PMC article. Review.
-
Transcriptome Characterization and Gene Changes Induced by Fusarium solani in Sweetpotato Roots.Genes (Basel). 2023 Apr 25;14(5):969. doi: 10.3390/genes14050969. Genes (Basel). 2023. PMID: 37239329 Free PMC article.
-
Genome-wide identification and expression analysis of the glutamate receptor gene family in sweet potato and its two diploid relatives.Front Plant Sci. 2023 Dec 21;14:1255805. doi: 10.3389/fpls.2023.1255805. eCollection 2023. Front Plant Sci. 2023. PMID: 38179475 Free PMC article.
-
Sweet Potato as a Key Crop for Food Security under the Conditions of Global Climate Change: A Review.Plants (Basel). 2023 Jun 30;12(13):2516. doi: 10.3390/plants12132516. Plants (Basel). 2023. PMID: 37447081 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

