Interaction of Amphipathic Peptide from Influenza Virus M1 Protein with Mitochondrial Cytochrome Oxidase
- PMID: 36835528
- PMCID: PMC9961948
- DOI: 10.3390/ijms24044119
Interaction of Amphipathic Peptide from Influenza Virus M1 Protein with Mitochondrial Cytochrome Oxidase
Abstract
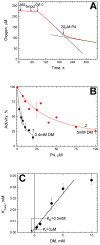
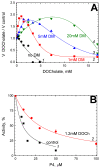
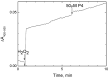
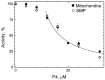
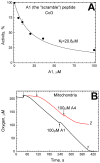
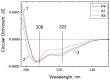
The Bile Acid Binding Site (BABS) of cytochrome oxidase (CcO) binds numerous amphipathic ligands. To determine which of the BABS-lining residues are critical for interaction, we used the peptide P4 and its derivatives A1-A4. P4 is composed of two flexibly bound modified α-helices from the M1 protein of the influenza virus, each containing a cholesterol-recognizing CRAC motif. The effect of the peptides on the activity of CcO was studied in solution and in membranes. The secondary structure of the peptides was examined by molecular dynamics, circular dichroism spectroscopy, and testing the ability to form membrane pores. P4 was found to suppress the oxidase but not the peroxidase activity of solubilized CcO. The Ki(app) is linearly dependent on the dodecyl-maltoside (DM) concentration, indicating that DM and P4 compete in a 1:1 ratio. The true Ki is 3 μM. The deoxycholate-induced increase in Ki(app) points to a competition between P4 and deoxycholate. A1 and A4 inhibit solubilized CcO with Ki(app)~20 μM at 1 mM DM. A2 and A3 hardly inhibit CcO either in solution or in membranes. The mitochondrial membrane-bound CcO retains sensitivity to P4 and A4 but acquires resistance to A1. We associate the inhibitory effect of P4 with its binding to BABS and dysfunction of the proton channel K. Trp residue is critical for inhibition. The resistance of the membrane-bound enzyme to inhibition may be due to the disordered secondary structure of the inhibitory peptide.
Keywords: Bile Acids Binding Site; amphipathic ligands; cell penetrating peptides; cytochrome oxidase; regulation; secondary structure; α-helix.
Conflict of interest statement
The authors declare no conflict of interest. The sponsors had no role in the design, execution, interpretation, or writing of the study.
Figures



Similar articles
-
Cholesterol Attenuates the Pore-Forming Capacity of CARC-Containing Amphipathic Peptides.Int J Mol Sci. 2025 Jan 10;26(2):533. doi: 10.3390/ijms26020533. Int J Mol Sci. 2025. PMID: 39859248 Free PMC article.
-
Mechanism of Inhibition of Cytochrome c Oxidase by Triton X-100.Biochemistry (Mosc). 2021 Jan;86(1):44-58. doi: 10.1134/S0006297921010053. Biochemistry (Mosc). 2021. PMID: 33705281
-
Direct Interaction of Mitochondrial Cytochrome c Oxidase with Thyroid Hormones: Evidence for Two Binding Sites.Cells. 2022 Mar 6;11(5):908. doi: 10.3390/cells11050908. Cells. 2022. PMID: 35269529 Free PMC article.
-
Role of conformational change and K-path ligands in controlling cytochrome c oxidase activity.Biochem Soc Trans. 2017 Oct 15;45(5):1087-1095. doi: 10.1042/BST20160138. Epub 2017 Aug 24. Biochem Soc Trans. 2017. PMID: 28842531 Free PMC article. Review.
-
Respiratory conservation of energy with dioxygen: cytochrome C oxidase.Met Ions Life Sci. 2015;15:89-130. doi: 10.1007/978-3-319-12415-5_4. Met Ions Life Sci. 2015. PMID: 25707467 Review.
Cited by
-
Mitochondrial Oxidative Phosphorylation in Viral Infections.Viruses. 2023 Dec 4;15(12):2380. doi: 10.3390/v15122380. Viruses. 2023. PMID: 38140621 Free PMC article. Review.
-
Interaction of Terminal Oxidases with Amphipathic Molecules.Int J Mol Sci. 2023 Mar 29;24(7):6428. doi: 10.3390/ijms24076428. Int J Mol Sci. 2023. PMID: 37047401 Free PMC article. Review.
-
Cholesterol Attenuates the Pore-Forming Capacity of CARC-Containing Amphipathic Peptides.Int J Mol Sci. 2025 Jan 10;26(2):533. doi: 10.3390/ijms26020533. Int J Mol Sci. 2025. PMID: 39859248 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

