Comparing the Efficacy of MALDI-TOF MS and Sequencing-Based Identification Techniques (Sanger and NGS) to Monitor the Microbial Community of Irrigation Water
- PMID: 36838251
- PMCID: PMC9960253
- DOI: 10.3390/microorganisms11020287
Comparing the Efficacy of MALDI-TOF MS and Sequencing-Based Identification Techniques (Sanger and NGS) to Monitor the Microbial Community of Irrigation Water
Abstract
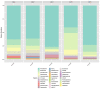
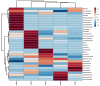
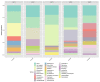
In order to intensify and guarantee the agricultural productivity and thereby to be able to feed the world's rapidly growing population, irrigation has become very important. In parallel, the limited water resources lead to an increase in usage of poorly characterized sources of water, which is directly linked to a higher prevalence of foodborne diseases. Therefore, analyzing the microorganisms or even the complete microbiome of irrigation water used for food production can prevent the growing numbers of such cases. In this study, we compared the efficacy of MALDI-TOF Mass spectrometry (MALDI TOF MS) identification to 16S rRNA gene Sanger sequencing of waterborne microorganisms. Furthermore, we analyzed the whole microbial community of irrigation water using high-throughput 16S rRNA gene amplicon sequencing. The identification results of MALDI-TOF MS and 16S rRNA gene Sanger sequencing were almost identical at species level (66.7%; 64.3%). Based on the applied cultivation techniques, Acinetobacter spp., Enterobacter spp., Pseudomonas spp., and Brevundimonas spp. were the most abundant cultivable genera. In addition, the uncultivable part of the microbiome was dominated by Proteobacteria followed by Actinobacteria, Bacteroidota, Patescibacteria, and Verrucomicrobiota. Our findings indicate that MALDI-TOF MS offers a fast, reliable identification method and can act as an alternative to 16S rRNA gene Sanger sequencing of isolates. Moreover, the results suggest that MALDI-TOF MS paired with 16S rRNA gene amplicon sequencing have the potential to support the routine monitoring of the microbiological quality of irrigation water.
Keywords: 16S rRNA gene amplicon sequencing; MALDI-TOF MS; Sanger sequencing; irrigation water; microbial monitoring.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Comparison of identification systems for psychrotrophic bacteria isolated from raw bovine milk.Int J Food Microbiol. 2014 Oct 17;189:26-38. doi: 10.1016/j.ijfoodmicro.2014.07.023. Epub 2014 Jul 26. Int J Food Microbiol. 2014. PMID: 25113043
-
Determination of the microbiological contamination in minced pork by culture dependent and 16S amplicon sequencing analysis.Int J Food Microbiol. 2019 Feb 2;290:27-35. doi: 10.1016/j.ijfoodmicro.2018.09.025. Epub 2018 Oct 2. Int J Food Microbiol. 2019. PMID: 30292676
-
Evaluation of Human Milk Microbiota by 16S rRNA Gene Next-Generation Sequencing (NGS) and Cultivation/MALDI-TOF Mass Spectrometry Identification.Front Microbiol. 2019 Nov 15;10:2612. doi: 10.3389/fmicb.2019.02612. eCollection 2019. Front Microbiol. 2019. PMID: 31803156 Free PMC article.
-
MALDI-TOF mass spectrometry: an emerging technology for microbial identification and diagnosis.Front Microbiol. 2015 Aug 5;6:791. doi: 10.3389/fmicb.2015.00791. eCollection 2015. Front Microbiol. 2015. PMID: 26300860 Free PMC article. Review.
-
Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass-Spectrometry (MALDI-TOF MS) Based Microbial Identifications: Challenges and Scopes for Microbial Ecologists.Front Microbiol. 2016 Aug 30;7:1359. doi: 10.3389/fmicb.2016.01359. eCollection 2016. Front Microbiol. 2016. PMID: 27625644 Free PMC article. Review.
Cited by
-
Integrating multi-wet laboratory diagnostics to study staphylococci in animals in Uganda.BMC Microbiol. 2024 Aug 10;24(1):298. doi: 10.1186/s12866-024-03442-x. BMC Microbiol. 2024. PMID: 39127665 Free PMC article.
-
Neisseria sicca Vertebral Osteomyelitis: A Case Report and Literature Review.J Clin Med. 2024 Nov 28;13(23):7241. doi: 10.3390/jcm13237241. J Clin Med. 2024. PMID: 39685699 Free PMC article.
-
Bacterial contamination of medical face mask wearing duration and the optimal wearing time.Front Cell Infect Microbiol. 2023 Oct 2;13:1231248. doi: 10.3389/fcimb.2023.1231248. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 37850052 Free PMC article.
-
Detection of Staphylococcus Isolates and Their Antimicrobial Resistance Profiles and Virulence Genes from Subclinical Mastitis Cattle Milk Using MALDI-TOF MS, PCR and Sequencing in Free State Province, South Africa.Animals (Basel). 2024 Jan 2;14(1):154. doi: 10.3390/ani14010154. Animals (Basel). 2024. PMID: 38200885 Free PMC article.
References
-
- Gu G., Luo Z., Cevallos-Cevallos J.M., Adams P., Vellidis G., Wright A., van Bruggen A.H. Factors affecting the occurrence of Escherichia coli O157 contamination in irrigation ponds on produce farms in the Suwannee River Watershed. Can. J. Microbiol. 2013;59:175–182. doi: 10.1139/cjm-2012-0599. - DOI - PubMed
-
- Uyttendaele M., Jaykus L.-A., Amoah P., Chiodini A., Cunliffe D., Jacxsens L., Holvoet K., Korsten L., Lau M., McClure P., et al. Microbial Hazards in Irrigation Water: Standards, Norms, and Testing to Manage Use of Water in Fresh Produce Primary Production. Compr. Rev. Food Sci. 2015;14:336–356. doi: 10.1111/1541-4337.12133. - DOI
-
- Falardeau J., Johnson R.P., Pagotto F., Wang S. Occurrence, characterization, and potential predictors of verotoxigenic Escherichia coli, Listeria monocytogenes, and Salmonella in surface water used for produce irrigation in the Lower Mainland of British Columbia, Canada. PloS ONE. 2017;27:9. doi: 10.1371/journal.pone.0185437. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

