Global landscape of SARS-CoV-2 mutations and conserved regions
- PMID: 36841805
- PMCID: PMC9958328
- DOI: 10.1186/s12967-023-03996-w
Global landscape of SARS-CoV-2 mutations and conserved regions
Abstract
Background: At the end of December 2019, a novel strain of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) disease (COVID-19) has been identified in Wuhan, a central city in China, and then spread to every corner of the globe. As of October 8, 2022, the total number of COVID-19 cases had reached over 621 million worldwide, with more than 6.56 million confirmed deaths. Since SARS-CoV-2 genome sequences change due to mutation and recombination, it is pivotal to surveil emerging variants and monitor changes for improving pandemic management.
Methods: 10,287,271 SARS-CoV-2 genome sequence samples were downloaded in FASTA format from the GISAID databases from February 24, 2020, to April 2022. Python programming language (version 3.8.0) software was utilized to process FASTA files to identify variants and sequence conservation. The NCBI RefSeq SARS-CoV-2 genome (accession no. NC_045512.2) was considered as the reference sequence.
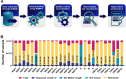
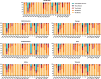
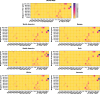
Results: Six mutations had more than 50% frequency in global SARS-CoV-2. These mutations include the P323L (99.3%) in NSP12, D614G (97.6) in S, the T492I (70.4) in NSP4, R203M (62.8%) in N, T60A (61.4%) in Orf9b, and P1228L (50.0%) in NSP3. In the SARS-CoV-2 genome, no mutation was observed in more than 90% of nsp11, nsp7, nsp10, nsp9, nsp8, and nsp16 regions. On the other hand, N, nsp3, S, nsp4, nsp12, and M had the maximum rate of mutations. In the S protein, the highest mutation frequency was observed in aa 508-635(0.77%) and aa 381-508 (0.43%). The highest frequency of mutation was observed in aa 66-88 (2.19%), aa 7-14, and aa 164-246 (2.92%) in M, E, and N proteins, respectively.
Conclusion: Therefore, monitoring SARS-CoV-2 proteomic changes and detecting hot spots mutations and conserved regions could be applied to improve the SARS-CoV-2 diagnostic efficiency and design safe and effective vaccines against emerging variants.
Keywords: Amino Acid; COVID-19; Emerging variants; Genome; SARS-CoV-2; Vaccines.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



Similar articles
-
Comparative Atlas of SARS-CoV-2 Substitution Mutations: A Focus on Iranian Strains Amidst Global Trends.Viruses. 2024 Aug 20;16(8):1331. doi: 10.3390/v16081331. Viruses. 2024. PMID: 39205305 Free PMC article.
-
Correlates of SARS-CoV-2 Variants on Deaths, Case Incidence and Case Fatality Ratio among the Continents for the Period of 1 December 2020 to 15 March 2021.Genes (Basel). 2021 Jul 12;12(7):1061. doi: 10.3390/genes12071061. Genes (Basel). 2021. PMID: 34356077 Free PMC article.
-
Coronavirus Genomes and Unique Mutations in Structural and Non-Structural Proteins in Pakistani SARS-CoV-2 Delta Variants during the Fourth Wave of the Pandemic.Genes (Basel). 2022 Mar 21;13(3):552. doi: 10.3390/genes13030552. Genes (Basel). 2022. PMID: 35328105 Free PMC article.
-
Characterization of SARS-CoV-2 different variants and related morbidity and mortality: a systematic review.Eur J Med Res. 2021 Jun 8;26(1):51. doi: 10.1186/s40001-021-00524-8. Eur J Med Res. 2021. PMID: 34103090 Free PMC article.
-
Variants of SARS CoV-2: mutations, transmissibility, virulence, drug resistance, and antibody/vaccine sensitivity.Front Biosci (Landmark Ed). 2022 Feb 14;27(2):65. doi: 10.31083/j.fbl2702065. Front Biosci (Landmark Ed). 2022. PMID: 35227008 Review.
Cited by
-
Factors influencing neutralizing antibody response to the original SARS-CoV-2 virus and the Omicron variant in a high vaccination coverage country, a population-based study.Vaccine X. 2023 Aug 26;15:100372. doi: 10.1016/j.jvacx.2023.100372. eCollection 2023 Dec. Vaccine X. 2023. PMID: 37693843 Free PMC article.
-
A Cullin 5-based complex serves as an essential modulator of ORF9b stability in SARS-CoV-2 replication.Signal Transduct Target Ther. 2024 Jun 28;9(1):159. doi: 10.1038/s41392-024-01874-5. Signal Transduct Target Ther. 2024. PMID: 38937432 Free PMC article.
-
A Bayesian inference method to estimate transmission trees with multiple introductions; applied to SARS-CoV-2 in Dutch mink farms.PLoS Comput Biol. 2023 Nov 27;19(11):e1010928. doi: 10.1371/journal.pcbi.1010928. eCollection 2023 Nov. PLoS Comput Biol. 2023. PMID: 38011266 Free PMC article.
-
Immunogenicity and Protective Efficacy of Baculovirus-Expressed SARS-CoV-2 Envelope Protein in Mice as a Universal Vaccine Candidate.Vaccines (Basel). 2024 Aug 28;12(9):977. doi: 10.3390/vaccines12090977. Vaccines (Basel). 2024. PMID: 39340009 Free PMC article.
-
Development and Evaluation of an In-House Real-Time RT-PCR Targeting nsp10 Gene for SARS-CoV-2 Detection.Int J Mol Sci. 2024 Mar 21;25(6):3552. doi: 10.3390/ijms25063552. Int J Mol Sci. 2024. PMID: 38542524 Free PMC article.
References
-
- WHO. Coronavirus disease (COVID-19) pandemic, 2022. https://www.who.int/emergencies/diseases/novelcoronavirus-2019.
-
- Aljindan RY, Al-Subaie AM, Al-Ohali AI, Kamaraj B. Investigation of nonsynonymous mutations in the spike protein of SARS-CoV-2 and its interaction with the ACE2 receptor by molecular docking and MM/GBSA approach. Comput Biol Med. 2021;135:104654. doi: 10.1016/j.compbiomed.2021.104654. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

