Design of a multi-epitope vaccine against six Nocardia species based on reverse vaccinology combined with immunoinformatics
- PMID: 36845087
- PMCID: PMC9952739
- DOI: 10.3389/fimmu.2023.1100188
Design of a multi-epitope vaccine against six Nocardia species based on reverse vaccinology combined with immunoinformatics
Abstract
Background: Nocardia genus, a complex group of species classified to be aerobic actinomycete, can lead to severe concurrent infection as well as disseminated infection, typically in immunocompromised patients. With the expansion of the susceptible population, the incidence of Nocardia has been gradually growing, accompanied by increased resistance of the pathogen to existing therapeutics. However, there is no effective vaccine against this pathogen yet. In this study, a multi-epitope vaccine was designed against the Nocardia infection using reverse vaccinology combined with immunoinformatics approaches.
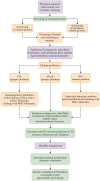
Methods: First, the proteomes of 6 Nocardia subspecies Nocardia subspecies (Nocardia farcinica, Nocardia cyriacigeorgica, Nocardia abscessus, Nocardia otitidiscaviarum, Nocardia brasiliensis and Nocardia nova) were download NCBI (National Center for Biotechnology Information) database on May 1st, 2022 for the target proteins selection. The essential, virulent-associated or resistant-associated, surface-exposed, antigenic, non-toxic, and non-homologous with the human proteome proteins were selected for epitope identification. The shortlisted T-cell and B-cell epitopes were fused with appropriate adjuvants and linkers to construct vaccines. The physicochemical properties of the designed vaccine were predicted using multiple online servers. The Molecular docking and molecular dynamics (MD) simulation were performed to understand the binding pattern and binding stability between the vaccine candidate and Toll-like receptors (TLRs). The immunogenicity of the designed vaccines was evaluated via immune simulation.
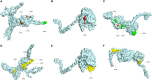
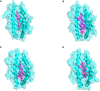
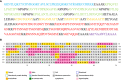
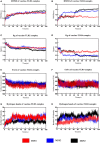
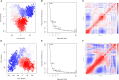
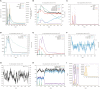
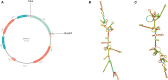
Results: 3 proteins that are essential, virulent-associated or resistant-associated, surface-exposed, antigenic, non-toxic, and non-homologous with the human proteome were selected from 218 complete proteome sequences of the 6 Nocardia subspecies epitope identification. After screening, only 4 cytotoxic T lymphocyte (CTL) epitopes, 6 helper T lymphocyte (HTL) epitopes, and 8 B cell epitopes that were antigenic, non-allergenic, and non-toxic were included in the final vaccine construct. The results of molecular docking and MD simulation showed that the vaccine candidate has a strong affinity for TLR2 and TLR4 of the host and the vaccine-TLR complexes were dynamically stable in the natural environment. The results of the immune simulation indicated that the designed vaccine had the potential to induce strong protective immune responses in the host. The codon optimization and cloned analysis showed that the vaccine was available for mass production.
Conclusion: The designed vaccine has the potential to stimulate long-lasting immunity in the host, but further studies are required to validate its safety and efficacy.
Keywords: Nocardia; immunoinformatics; molecular docking; molecular dynamics (MD) simulation; multi-epitope vaccine; reverse vaccinology (RV).
Copyright © 2023 Zhu, Tan, Li, Ma, Wen, Yang, Rao, Zhang, Peng, Cui, Chen and Pan.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Prioritization of potential vaccine candidates and designing a multiepitope-based subunit vaccine against multidrug-resistant Salmonella Typhi str. CT18: A subtractive proteomics and immunoinformatics approach.Microb Pathog. 2021 Oct;159:105150. doi: 10.1016/j.micpath.2021.105150. Epub 2021 Aug 20. Microb Pathog. 2021. PMID: 34425197
-
Development of a novel multi‑epitope vaccine against the pathogenic human polyomavirus V6/7 using reverse vaccinology.BMC Infect Dis. 2024 Feb 9;24(1):177. doi: 10.1186/s12879-024-09046-0. BMC Infect Dis. 2024. PMID: 38336665 Free PMC article.
-
Immunoinformatics design of a novel epitope-based vaccine candidate against dengue virus.Sci Rep. 2021 Oct 5;11(1):19707. doi: 10.1038/s41598-021-99227-7. Sci Rep. 2021. PMID: 34611250 Free PMC article.
-
Reverse engineering protection: A comprehensive survey of reverse vaccinology-based vaccines targeting viral pathogens.Vaccine. 2024 Apr 11;42(10):2503-2518. doi: 10.1016/j.vaccine.2024.02.087. Epub 2024 Mar 23. Vaccine. 2024. PMID: 38523003 Review.
-
Immunogenic multi-epitope-based vaccine development to combat cyclosporiasis of immunocompromised patients applying computational biology method.Exp Parasitol. 2023 May;248:108497. doi: 10.1016/j.exppara.2023.108497. Epub 2023 Mar 10. Exp Parasitol. 2023. PMID: 36906252 Review.
Cited by
-
A computational approach to developing a multi-epitope vaccine for combating Pseudomonas aeruginosa-induced pneumonia and sepsis.Brief Bioinform. 2024 Jul 25;25(5):bbae401. doi: 10.1093/bib/bbae401. Brief Bioinform. 2024. PMID: 39133098 Free PMC article.
-
Immunoinformatic evaluation for the development of a potent multi-epitope vaccine against bacterial vaginosis caused by Gardnerella vaginalis.PLoS One. 2025 Feb 27;20(2):e0316699. doi: 10.1371/journal.pone.0316699. eCollection 2025. PLoS One. 2025. PMID: 40014550 Free PMC article.
-
The Screening of the Protective Antigens of Aeromonas hydrophila Using the Reverse Vaccinology Approach: Potential Candidates for Subunit Vaccine Development.Vaccines (Basel). 2023 Jul 21;11(7):1266. doi: 10.3390/vaccines11071266. Vaccines (Basel). 2023. PMID: 37515081 Free PMC article.
-
Design and in silico analysis of a novel peptide-based multiepitope vaccine against glioblastoma multiforme by targeting tumor-associated macrophage.Heliyon. 2024 Nov 28;10(24):e40774. doi: 10.1016/j.heliyon.2024.e40774. eCollection 2024 Dec 30. Heliyon. 2024. PMID: 39759328 Free PMC article.
-
Innovations, Challenges, and Future Prospects for Combination Vaccines Against Human Infections.Vaccines (Basel). 2025 Mar 21;13(4):335. doi: 10.3390/vaccines13040335. Vaccines (Basel). 2025. PMID: 40333234 Free PMC article. Review.
References
-
- Hamdi AM, Fida M, Deml SM, Abu Saleh OM, Wengenack NL. Retrospective analysis of antimicrobial susceptibility profiles of nocardia species from a tertiary hospital and reference laboratory, 2011 to 2017. Antimicrob Agents Chemother (2020) 64(3):20200221. doi: 10.1128/aac.01868-19 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

