Specialized DNA Structures Act as Genomic Beacons for Integration by Evolutionarily Diverse Retroviruses
- PMID: 36851678
- PMCID: PMC9962126
- DOI: 10.3390/v15020465
Specialized DNA Structures Act as Genomic Beacons for Integration by Evolutionarily Diverse Retroviruses
Abstract
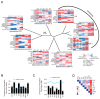
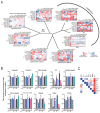
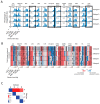
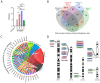
Retroviral integration site targeting is not random and plays a critical role in expression and long-term survival of the integrated provirus. To better understand the genomic environment surrounding retroviral integration sites, we performed a meta-analysis of previously published integration site data from evolutionarily diverse retroviruses, including new experimental data from HIV-1 subtypes A, B, C and D. We show here that evolutionarily divergent retroviruses exhibit distinct integration site profiles with strong preferences for integration near non-canonical B-form DNA (non-B DNA). We also show that in vivo-derived HIV-1 integration sites are significantly more enriched in transcriptionally silent regions and transcription-silencing non-B DNA features of the genome compared to in vitro-derived HIV-1 integration sites. Integration sites from individuals infected with HIV-1 subtype A, B, C or D viruses exhibited different preferences for common genomic and non-B DNA features. In addition, we identified several integration site hotspots shared between different HIV-1 subtypes, all of which were located in the non-B DNA feature slipped DNA. Together, these data show that although evolutionarily divergent retroviruses exhibit distinct integration site profiles, they all target non-B DNA for integration. These findings provide new insight into how retroviruses integrate into genomes for long-term survival.
Keywords: HIV; genome; integration; integration hotspots; non-B DNA; retroviruses; slipped DNA.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


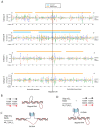
References
-
- Daniel R., Greger J.G., Katz R.A., Taganov K.D., Wu X., Kappes J.C., Skalka A.M. Evidence That Stable Retroviral Transduction and Cell Survival Following DNA Integration Depend on Components of the Nonhomologous End Joining Repair Pathway. J. Virol. 2004;78:8573–8581. doi: 10.1128/JVI.78.16.8573-8581.2004. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

