Circulating Interleukin-8 Dynamics Parallels Disease Course and Is Linked to Clinical Outcomes in Severe COVID-19
- PMID: 36851762
- PMCID: PMC9960352
- DOI: 10.3390/v15020549
Circulating Interleukin-8 Dynamics Parallels Disease Course and Is Linked to Clinical Outcomes in Severe COVID-19
Abstract
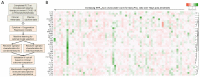
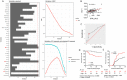
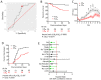
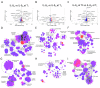
Severe COVID-19 frequently features a systemic deluge of cytokines. Circulating cytokines that can stratify risks are useful for more effective triage and management. Here, we ran a machine-learning algorithm on a dataset of 36 plasma cytokines in a cohort of severe COVID-19 to identify cytokine/s useful for describing the dynamic clinical state in multiple regression analysis. We performed RNA-sequencing of circulating blood cells collected at different time-points. From a Bayesian Information Criterion analysis, a combination of interleukin-8 (IL-8), Eotaxin, and Interferon-γ (IFNγ) was found to be significantly linked to blood oxygenation over seven days. Individually testing the cytokines in receiver operator characteristics analyses identified IL-8 as a strong stratifier for clinical outcomes. Circulating IL-8 dynamics paralleled disease course. We also revealed key transitions in immune transcriptome in patients stratified for circulating IL-8 at three time-points. The study identifies plasma IL-8 as a key pathogenic cytokine linking systemic hyper-inflammation to the clinical outcomes in COVID-19.
Keywords: COVID-19; IL-8; biomarker; cytokines; machine learning; survival.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures



References
-
- Edara V.V., Pinsky B.A., Suthar M.S., Lai L., Davis-Gardner M.E., Floyd K., Flowers M.W., Wrammert J., Hussaini L., Ciric C.R., et al. Infection and Vaccine-Induced Neutralizing-Antibody Responses to the SARS-CoV-2 B.1.617 Variants. N. Engl. J. Med. 2021;385:664–666. doi: 10.1056/NEJMc2107799. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

