Interpretable meta-learning of multi-omics data for survival analysis and pathway enrichment
- PMID: 36864611
- PMCID: PMC10079355
- DOI: 10.1093/bioinformatics/btad113
Interpretable meta-learning of multi-omics data for survival analysis and pathway enrichment
Abstract
Motivation: Despite the success of recent machine learning algorithms' applications to survival analysis, their black-box nature hinders interpretability, which is arguably the most important aspect. Similarly, multi-omics data integration for survival analysis is often constrained by the underlying relationships and correlations that are rarely well understood. The goal of this work is to alleviate the interpretability problem in machine learning approaches for survival analysis and also demonstrate how multi-omics data integration improves survival analysis and pathway enrichment. We use meta-learning, a machine-learning algorithm that is trained on a variety of related datasets and allows quick adaptations to new tasks, to perform survival analysis and pathway enrichment on pan-cancer datasets. In recent machine learning research, meta-learning has been effectively used for knowledge transfer among multiple related datasets.
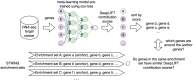
Results: We use meta-learning with Cox hazard loss to show that the integration of TCGA pan-cancer data increases the performance of survival analysis. We also apply advanced model interpretability method called DeepLIFT (Deep Learning Important FeaTures) to show different sets of enriched pathways for multi-omics and transcriptomics data. Our results show that multi-omics cancer survival analysis enhances performance compared with using transcriptomics or clinical data alone. Additionally, we show a correlation between variable importance assignment from DeepLIFT and gene coenrichment, suggesting that genes with higher and similar contribution scores are more likely to be enriched together in the same enrichment sets.
Availability and implementation: https://github.com/berkuva/TCGA-omics-integration.
© The Author(s) 2023. Published by Oxford University Press.
Figures

Similar articles
-
MMOSurv: meta-learning for few-shot survival analysis with multi-omics data.Bioinformatics. 2024 Dec 26;41(1):btae684. doi: 10.1093/bioinformatics/btae684. Bioinformatics. 2024. PMID: 39563482 Free PMC article.
-
Cancer subtype identification by multi-omics clustering based on interpretable feature and latent subspace learning.Methods. 2024 Nov;231:144-153. doi: 10.1016/j.ymeth.2024.09.014. Epub 2024 Sep 24. Methods. 2024. PMID: 39326482
-
A fair experimental comparison of neural network architectures for latent representations of multi-omics for drug response prediction.BMC Bioinformatics. 2023 Feb 14;24(1):45. doi: 10.1186/s12859-023-05166-7. BMC Bioinformatics. 2023. PMID: 36788531 Free PMC article.
-
A critical review of machine-learning for "multi-omics" marine metabolite datasets.Comput Biol Med. 2023 Oct;165:107425. doi: 10.1016/j.compbiomed.2023.107425. Epub 2023 Aug 29. Comput Biol Med. 2023. PMID: 37696182 Review.
-
Interpretable machine learning methods for predictions in systems biology from omics data.Front Mol Biosci. 2022 Oct 17;9:926623. doi: 10.3389/fmolb.2022.926623. eCollection 2022. Front Mol Biosci. 2022. PMID: 36387282 Free PMC article. Review.
Cited by
-
Integrative analysis of cancer multimodality data identifying COPS5 as a novel biomarker of diffuse large B-cell lymphoma.Front Genet. 2024 Jun 21;15:1407765. doi: 10.3389/fgene.2024.1407765. eCollection 2024. Front Genet. 2024. PMID: 38974382 Free PMC article.
-
Federated transfer learning with differential privacy for multi-omics survival analysis.Brief Bioinform. 2025 Mar 4;26(2):bbaf166. doi: 10.1093/bib/bbaf166. Brief Bioinform. 2025. PMID: 40230038 Free PMC article.
-
A review of multi-omics data integration through deep learning approaches for disease diagnosis, prognosis, and treatment.Front Genet. 2023 Jul 20;14:1199087. doi: 10.3389/fgene.2023.1199087. eCollection 2023. Front Genet. 2023. PMID: 37547471 Free PMC article. Review.
-
IBPGNET: lung adenocarcinoma recurrence prediction based on neural network interpretability.Brief Bioinform. 2024 Mar 27;25(3):bbae080. doi: 10.1093/bib/bbae080. Brief Bioinform. 2024. PMID: 38557672 Free PMC article.
-
Single-cell transcriptome analysis revealed heterogeneity in glycolysis and identified IGF2 as a therapeutic target for ovarian cancer subtypes.BMC Cancer. 2024 Jul 31;24(1):926. doi: 10.1186/s12885-024-12688-7. BMC Cancer. 2024. PMID: 39085784 Free PMC article.
References
-
- Alipanahi B, Delong A, Weirauch MT. et al. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat Biotechnol 2015;33:831–8. - PubMed
-
- Bai Y, Wang G, Fu W. et al. Circulating essential metals and lung cancer: risk assessment and potential molecular effects. Environ Int 2019;127:685–93. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

