JMJD6 Shapes a Pro-tumor Microenvironment via ANXA1-Dependent Macrophage Polarization in Breast Cancer
- PMID: 36867680
- PMCID: PMC10233359
- DOI: 10.1158/1541-7786.MCR-22-0370
JMJD6 Shapes a Pro-tumor Microenvironment via ANXA1-Dependent Macrophage Polarization in Breast Cancer
Abstract
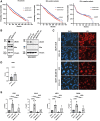
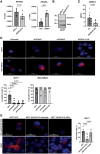
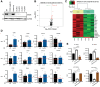
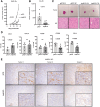
Breast cancer is the most common type of cancer in women worldwide, with the luminal subtype being the most widespread. Although characterized by better prognosis compared with other subtypes, luminal breast cancer is still considered a threatening disease due to therapy resistance, which occurs via both cell- and non-cell-autonomous mechanisms. Jumonji domain-containing 6, arginine demethylase and lysine hydroxylase (JMJD6) is endowed with a negative prognostic value in luminal breast cancer and, via its epigenetic activity, it is known to regulate many intrinsic cancer cell pathways. So far, the effect of JMJD6 in molding the surrounding microenvironment has not been explored.Here, we describe a novel function of JMJD6 showing that its genetic inhibition in breast cancer cells suppresses lipid droplet formation and ANXA1 expression, via estrogen receptor alpha and PPARα modulation. Reduction of intracellular ANXA1 results in decreased release in the tumor microenvironment (TME), ultimately preventing M2-type macrophage polarization and tumor aggressiveness.
Implications: Our findings identify JMJD6 as a determinant of breast cancer aggressiveness and provide the rationale for the development of inhibitory molecules to reduce disease progression also through the remodeling of TME composition.
©2023 The Authors; Published by the American Association for Cancer Research.
Figures


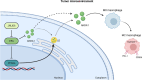
References
-
- Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021;71:209–49. - PubMed
-
- DeSantis CE, Ma J, Gaudet MM, Newman LA, Miller KD, Goding Sauer A, et al. Breast cancer statistics, 2019. CA Cancer J Clin 2019;69:438–51. - PubMed
-
- Carter CL, Allen C, Henson DE. Relation of tumor size, lymph node status, and survival in 24,740 breast cancer cases. Cancer 1989;63:181–7. - PubMed
-
- Andre F, Pusztai L. Molecular classification of breast cancer: implications for selection of adjuvant chemotherapy. Nat Clin Pract Oncol 2006;3:621–32. - PubMed
-
- Schnitt SJ. Classification and prognosis of invasive breast cancer: from morphology to molecular taxonomy. Mod Pathol 2010;23:S60–4. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

