The revised reference genome of the leopard gecko (Eublepharis macularius) provides insight into the considerations of genome phasing and assembly
- PMID: 36869788
- PMCID: PMC10445513
- DOI: 10.1093/jhered/esad016
The revised reference genome of the leopard gecko (Eublepharis macularius) provides insight into the considerations of genome phasing and assembly
Abstract
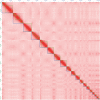
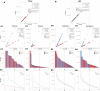
Genomic resources across squamate reptiles (lizards and snakes) have lagged behind other vertebrate systems and high-quality reference genomes remain scarce. Of the 23 chromosome-scale reference genomes across the order, only 12 of the ~60 squamate families are represented. Within geckos (infraorder Gekkota), a species-rich clade of lizards, chromosome-level genomes are exceptionally sparse representing only two of the seven extant families. Using the latest advances in genome sequencing and assembly methods, we generated one of the highest-quality squamate genomes to date for the leopard gecko, Eublepharis macularius (Eublepharidae). We compared this assembly to the previous, short-read only, E. macularius reference genome published in 2016 and examined potential factors within the assembly influencing contiguity of genome assemblies using PacBio HiFi data. Briefly, the read N50 of the PacBio HiFi reads generated for this study was equal to the contig N50 of the previous E. macularius reference genome at 20.4 kilobases. The HiFi reads were assembled into a total of 132 contigs, which was further scaffolded using HiC data into 75 total sequences representing all 19 chromosomes. We identified 9 of the 19 chromosomal scaffolds were assembled as a near-single contig, whereas the other 10 chromosomes were each scaffolded together from multiple contigs. We qualitatively identified that the percent repeat content within a chromosome broadly affects its assembly contiguity prior to scaffolding. This genome assembly signifies a new age for squamate genomics where high-quality reference genomes rivaling some of the best vertebrate genome assemblies can be generated for a fraction of previous cost estimates. This new E. macularius reference assembly is available on NCBI at JAOPLA010000000.
Keywords: emerging model system; evolution; gekkota; genomics; phasing.
© The Author(s) 2023. Published by Oxford University Press on behalf of The American Genetic Association. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Figures

Update of
-
The revised reference genome of the leopard gecko ( Eublepharis macularius ) provides insight into the considerations of genome phasing and assembly.bioRxiv [Preprint]. 2023 Feb 13:2023.01.20.523807. doi: 10.1101/2023.01.20.523807. bioRxiv. 2023. Update in: J Hered. 2023 Aug 23;114(5):513-520. doi: 10.1093/jhered/esad016. PMID: 36712019 Free PMC article. Updated. Preprint.
References
-
- Adler D, Kelly ST, Elliott T, Adamson J.. vioplot: violin plot. R package version 0.4.0. 2022. https://github.com/TomKellyGenetics/vioplot.
-
- Agarwal I, Bauer AM, Gamble T, Giri VB, Jablonski D, Khandekar A, Mohapatra PP, Masroor R, Mishra A, Ramakrishnan U.. The evolutionary history of an accidental model organism, the leopard gecko Eublepharis macularius (Squamata: Eublepharidae). Mol Phylogenet Evol. 2022:168:107414. - PubMed
-
- Bauer AM. Geckos: the animal answer guide. Baltimore, MD: Johns Hopkins University Press; 2013.
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources

