APOE modulates microglial immunometabolism in response to age, amyloid pathology, and inflammatory challenge
- PMID: 36871219
- PMCID: PMC10117631
- DOI: 10.1016/j.celrep.2023.112196
APOE modulates microglial immunometabolism in response to age, amyloid pathology, and inflammatory challenge
Abstract
The E4 allele of Apolipoprotein E (APOE) is associated with both metabolic dysfunction and a heightened pro-inflammatory response: two findings that may be intrinsically linked through the concept of immunometabolism. Here, we combined bulk, single-cell, and spatial transcriptomics with cell-specific and spatially resolved metabolic analyses in mice expressing human APOE to systematically address the role of APOE across age, neuroinflammation, and AD pathology. RNA sequencing (RNA-seq) highlighted immunometabolic changes across the APOE4 glial transcriptome, specifically in subsets of metabolically distinct microglia enriched in the E4 brain during aging or following an inflammatory challenge. E4 microglia display increased Hif1α expression and a disrupted tricarboxylic acid (TCA) cycle and are inherently pro-glycolytic, while spatial transcriptomics and mass spectrometry imaging highlight an E4-specific response to amyloid that is characterized by widespread alterations in lipid metabolism. Taken together, our findings emphasize a central role for APOE in regulating microglial immunometabolism and provide valuable, interactive resources for discovery and validation research.
Keywords: APOE; Apolipoprotein E; CP: Neuroscience; DAM; LPS; aging; amyloid; immunometabolism; microglia; scRNA-seq; spatial transcriptomics.
Copyright © 2023 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



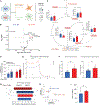



References
-
- Jansen IE, Savage JE, Watanabe K, Bryois J, Williams DM, Steinberg S, Sealock J, Karlsson IK, Hägg S, Athanasiu L, et al. (2019). Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet 51, 404–413. 10.1038/s41588-018-0311-9. - DOI - PMC - PubMed
-
- Kunkle BW, Grenier-Boley B, Sims R, Bis JC, Damotte V, Naj AC, Boland A, Vronskaya M, van der Lee SJ, Amlie-Wolf A, et al. (2019). Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Ab, tau, immunity and lipid processing. Nat. Genet 51, 414–430. 10.1038/s41588-019-0358-2. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

