Echinacea purpurea-derived homogeneous polysaccharide exerts anti-tumor efficacy via facilitating M1 macrophage polarization
- PMID: 36873268
- PMCID: PMC9974447
- DOI: 10.1016/j.xinn.2023.100391
Echinacea purpurea-derived homogeneous polysaccharide exerts anti-tumor efficacy via facilitating M1 macrophage polarization
Abstract
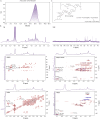
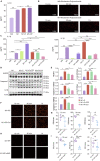
Echinacea purpurea modulates tumor progression, but the underlying mechanism is poorly defined. We isolated and purified a novel homogeneous polysaccharide from E. purpurea (EPPA), which was shown to be an arabinogalactan with a mean molecular mass (Mr) of 3.8 × 104 Da and with α- (1 → 5) -L-Arabinan as the backbone and α-L-Araf-(1→, →6)-β-D-Galp-(1→, and →4)-α-D-GalpA-(1→ as the side chains. Interestingly, oral administration of EPPA suppresses tumor progression in vivo and shapes the immune cell profile (e.g., facilitating M1 macrophages) in tumor microenvironment by single-cell RNA sequencing (scRNA-seq) analysis. More importantly, EPPA activates the inflammasome through a phagocytosis-dependent mechanism and rewires transcriptomic and metabolic profile, thereby potentiating M1 macrophage polarization. Collectively, we propose that EPPA supplementation could function as an adjuvant therapeutic strategy for tumor suppression.
© 2023 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
An arabinogalactan isolated from Pollen Typhae induces the apoptosis of RKO cells by promoting macrophage polarization.Carbohydr Polym. 2023 Jan 1;299:120216. doi: 10.1016/j.carbpol.2022.120216. Epub 2022 Oct 14. Carbohydr Polym. 2023. PMID: 36876818
-
Synthesis of the alpha-L-Araf-(1-->2)-beta-D-Galp-(1-->6)-beta-D-Galp-(1-->6)-[alpha-L-Araf-(1-->2)]-beta-D-Galp-(1-->6)-D-Gal hexasaccharide as a possible repeating unit of the cell-cultured exudates of Echinacea purpurea arabinogalactan.Carbohydr Res. 2001 Nov 8;336(2):107-15. doi: 10.1016/s0008-6215(01)00232-4. Carbohydr Res. 2001. PMID: 11689181
-
Structural Characterization and Macrophage Polarization-Modulating Activity of a Novel Polysaccharide from Large Yellow Tea.J Agric Food Chem. 2022 Oct 5;70(39):12565-12576. doi: 10.1021/acs.jafc.2c05593. Epub 2022 Sep 26. J Agric Food Chem. 2022. PMID: 36154025
-
Echinacea purpurea Extract Polarizes M1 Macrophages in Murine Bone Marrow-Derived Macrophages Through the Activation of JNK.J Cell Biochem. 2017 Sep;118(9):2664-2671. doi: 10.1002/jcb.25875. Epub 2017 May 15. J Cell Biochem. 2017. PMID: 28067413
-
Macrophage activation by the polysaccharide arabinogalactan isolated from plant cell cultures of Echinacea purpurea.J Natl Cancer Inst. 1989 May 3;81(9):669-75. doi: 10.1093/jnci/81.9.669. J Natl Cancer Inst. 1989. PMID: 2785214
Cited by
-
Echinacea: Bioactive Compounds and Agronomy.Plants (Basel). 2024 Apr 29;13(9):1235. doi: 10.3390/plants13091235. Plants (Basel). 2024. PMID: 38732450 Free PMC article. Review.
-
Steady-State Delivery and Chemical Modification of Food Nutrients to Improve Cancer Intervention Ability.Foods. 2024 Apr 28;13(9):1363. doi: 10.3390/foods13091363. Foods. 2024. PMID: 38731734 Free PMC article. Review.
-
Zi Shen Wan Fang repaired blood-brain barrier integrity in diabetic cognitive impairment mice via preventing cerebrovascular cells senescence.Chin Med. 2024 Dec 19;19(1):169. doi: 10.1186/s13020-024-01041-6. Chin Med. 2024. PMID: 39696612 Free PMC article.
-
Piriformospora indica culture filtrate and cell extract induce chicoric acid production in Echinacea purpurea hairy roots.PLoS One. 2025 Jun 17;20(6):e0323961. doi: 10.1371/journal.pone.0323961. eCollection 2025. PLoS One. 2025. PMID: 40526612 Free PMC article.
-
Echinacea purpurea (L.) Moench Polysaccharide Alleviates DSS-Induced Colitis in Rats by Restoring Th17/Treg Balance and Regulating Intestinal Flora.Foods. 2023 Nov 25;12(23):4265. doi: 10.3390/foods12234265. Foods. 2023. PMID: 38231750 Free PMC article.
References
-
- Wang Y., Wang Y., Ren Y., et al. Metabolic modulation of immune checkpoints and novel therapeutic strategies in cancer. Semin. Cancer Biol. 2022;86:542–565. - PubMed
-
- Nicolas A.M., Pesic M., Engel E., et al. Inflammatory fibroblasts mediate resistance to neoadjuvant therapy in rectal cancer. Cancer Cell. 2022;40:168–184.e13. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

