Nucleic acid nanostructures for in vivo applications: The influence of morphology on biological fate
- PMID: 36874908
- PMCID: PMC9869343
- DOI: 10.1063/5.0121820
Nucleic acid nanostructures for in vivo applications: The influence of morphology on biological fate
Abstract
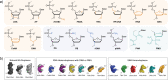
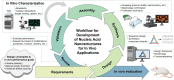
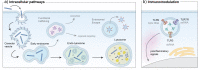
The development of programmable biomaterials for use in nanofabrication represents a major advance for the future of biomedicine and diagnostics. Recent advances in structural nanotechnology using nucleic acids have resulted in dramatic progress in our understanding of nucleic acid-based nanostructures (NANs) for use in biological applications. As the NANs become more architecturally and functionally diverse to accommodate introduction into living systems, there is a need to understand how critical design features can be controlled to impart desired performance in vivo. In this review, we survey the range of nucleic acid materials utilized as structural building blocks (DNA, RNA, and xenonucleic acids), the diversity of geometries for nanofabrication, and the strategies to functionalize these complexes. We include an assessment of the available and emerging characterization tools used to evaluate the physical, mechanical, physiochemical, and biological properties of NANs in vitro. Finally, the current understanding of the obstacles encountered along the in vivo journey is contextualized to demonstrate how morphological features of NANs influence their biological fates. We envision that this summary will aid researchers in the designing novel NAN morphologies, guide characterization efforts, and design of experiments and spark interdisciplinary collaborations to fuel advancements in programmable platforms for biological applications.
© 2023 Author(s).
Figures








References
-
- Seeman N. C. and Sleiman H. F., “ DNA nanotechnology,” Nat. Rev. Mater. 3(1), 17068 (2017). 10.1038/natrevmats.2017.68 - DOI
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
