SARS-CoV-2-encoded small RNAs are able to repress the host expression of SERINC5 to facilitate viral replication
- PMID: 36876111
- PMCID: PMC9978209
- DOI: 10.3389/fmicb.2023.1066493
SARS-CoV-2-encoded small RNAs are able to repress the host expression of SERINC5 to facilitate viral replication
Abstract
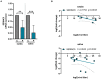
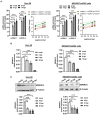
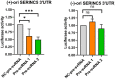
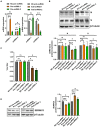
Serine incorporator protein 5 (SERINC5) is a key innate immunity factor that operates in the cell to restrict the infectivity of certain viruses. Different viruses have developed strategies to antagonize SERINC5 function but, how SERINC5 is controlled during viral infection is poorly understood. Here, we report that SERINC5 levels are reduced in COVID-19 patients during the infection by SARS-CoV-2 and, since no viral protein capable of repressing the expression of SERINC5 has been identified, we hypothesized that SARS-CoV-2 non-coding small viral RNAs (svRNAs) could be responsible for this repression. Two newly identified svRNAs with predicted binding sites in the 3'-untranslated region (3'-UTR) of the SERINC5 gene were characterized and we found that the expression of both svRNAs during the infection was not dependent on the miRNA pathway proteins Dicer and Argonaute-2. By using svRNAs mimic oligonucleotides, we demonstrated that both viral svRNAs can bind the 3'UTR of SERINC5 mRNA, reducing SERINC5 expression in vitro. Moreover, we found that an anti-svRNA treatment to Vero E6 cells before SARS-CoV-2 infection recovered the levels of SERINC5 and reduced the levels of N and S viral proteins. Finally, we showed that SERINC5 positively controls the levels of Mitochondrial Antiviral Signalling (MAVS) protein in Vero E6. These results highlight the therapeutic potential of targeting svRNAs based on their action on key proteins of the innate immune response during SARS-CoV-2 viral infection.
Keywords: MAVS; SARS-CoV-2; SERINC5; innate immune response; viral miRNAs.
Copyright © 2023 Meseguer, Rubio, Lainez, Pérez-Benavente, Pérez-Moraga, Romera-Giner, García-García, Martinez-Macias, Cremades, Iborra, Candelas-Rivera, Almazan and Esplugues.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures





References
-
- Ahi Y. S., Zhang S., Thappeta Y., Denman A., Feizpour A., Gummuluru S., et al. . (2016). Functional interplay between murine leukemia virus Glycogag, SERINC5, and surface glycoprotein governs virus entry, with opposite effects on Gammaretroviral and ebolavirus glycoproteins. MBio 7:e01985. doi: 10.1128/mBio.01985-16, PMID: - DOI - PMC - PubMed
-
- Chande A., Cuccurullo E. C., Rosa A., Ziglio S., Carpenter S., Pizzato M. (2016). S2 from equine infectious anemia virus is an infectivity factor which counteracts the retroviral inhibitors SERINC5 and SERINC3. Proc. Natl. Acad. Sci. U. S. A. 113, 13197–13202. doi: 10.1073/pnas.1612044113, PMID: - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

