Primary cilia sense glutamine availability and respond via asparagine synthetase
- PMID: 36879119
- PMCID: PMC10042734
- DOI: 10.1038/s42255-023-00754-6
Primary cilia sense glutamine availability and respond via asparagine synthetase
Abstract
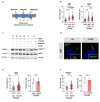
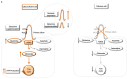
Depriving cells of nutrients triggers an energetic crisis, which is resolved by metabolic rewiring and organelle reorganization. Primary cilia are microtubule-based organelles at the cell surface, capable of integrating multiple metabolic and signalling cues, but their precise sensory function is not fully understood. Here we show that primary cilia respond to nutrient availability and adjust their length via glutamine-mediated anaplerosis facilitated by asparagine synthetase (ASNS). Nutrient deprivation causes cilia elongation, mediated by reduced mitochondrial function, ATP availability and AMPK activation independently of mTORC1. Of note, glutamine removal and replenishment is necessary and sufficient to induce ciliary elongation or retraction, respectively, under nutrient stress conditions both in vivo and in vitro by restoring mitochondrial anaplerosis via ASNS-dependent glutamate generation. Ift88-mutant cells lacking cilia show reduced glutamine-dependent mitochondrial anaplerosis during metabolic stress, due to reduced expression and activity of ASNS at the base of cilia. Our data indicate a role for cilia in responding to, and possibly sensing, cellular glutamine levels via ASNS during metabolic stress.
© 2023. The Author(s).
Conflict of interest statement
A.B. and M.C. are co-inventors on a patent for inhibition of glycolysis in PKD (patent family PCT/EP2013/064036). A.B., C.P. and M.C. are co-inventors on a patent for inhibiting ASNS in PKD (patent family PCT/EP2019/073628). All other authors declare no competing interests.
Figures









References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

