Single-cell profiling reveals pathogenic role and differentiation trajectory of granzyme K+CD8+ T cells in primary Sjögren's syndrome
- PMID: 36881472
- PMCID: PMC10243796
- DOI: 10.1172/jci.insight.167490
Single-cell profiling reveals pathogenic role and differentiation trajectory of granzyme K+CD8+ T cells in primary Sjögren's syndrome
Abstract
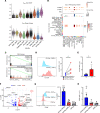
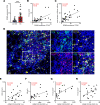
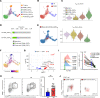
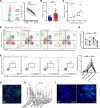
Primary Sjögren's syndrome (pSS) is a systemic autoimmune inflammatory disease mainly defined by T cell-dominated destruction of exocrine glands. Currently, CD8+ T cells are thought to be involved in the pathogenesis of pSS. However, the single-cell immune profiling of pSS and molecular signatures of pathogenic CD8+ T cells have not been well elucidated. Our multiomics investigation showed that both T cells and B cells, especially CD8+ T cells, were undergoing significant clonal expansion in pSS patients. TCR clonality analysis revealed that peripheral blood granzyme K+ (GZMK+) CXCR6+CD8+ T cells had higher a proportion of clones shared with CD69+CD103-CD8+ tissue-resident memory T (Trm) cells in labial glands in pSS. CD69+CD103-CD8+ Trm cells featured by high expression of GZMK were more active and cytotoxic in pSS compared with their CD103+ counterparts. Peripheral blood GZMK+CXCR6+CD8+ T cells with higher CD122 expression were increased and harbored a gene signature similar to Trm cells in pSS. Consistently, IL-15 was significantly elevated in pSS plasma and showed the capacity to promote differentiation of CD8+ T cells into GZMK+CXCR6+CD8+ T cells in a STAT5-dependent manner. In summary, we depicted the immune profile of pSS and further conducted comprehensive bioinformatics analysis and in vitro experimental investigations to characterize the pathogenic role and differentiation trajectory of CD8+ Trm cells in pSS.
Keywords: Autoimmune diseases; Autoimmunity; Immunology; Rheumatology; T cells.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

