Macrophage fumarate hydratase restrains mtRNA-mediated interferon production
- PMID: 36890227
- PMCID: PMC10411300
- DOI: 10.1038/s41586-023-05720-6
Macrophage fumarate hydratase restrains mtRNA-mediated interferon production
Abstract
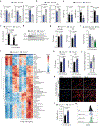
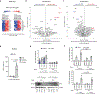
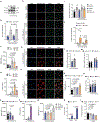
Metabolic rewiring underlies the effector functions of macrophages1-3, but the mechanisms involved remain incompletely defined. Here, using unbiased metabolomics and stable isotope-assisted tracing, we show that an inflammatory aspartate-argininosuccinate shunt is induced following lipopolysaccharide stimulation. The shunt, supported by increased argininosuccinate synthase (ASS1) expression, also leads to increased cytosolic fumarate levels and fumarate-mediated protein succination. Pharmacological inhibition and genetic ablation of the tricarboxylic acid cycle enzyme fumarate hydratase (FH) further increases intracellular fumarate levels. Mitochondrial respiration is also suppressed and mitochondrial membrane potential increased. RNA sequencing and proteomics analyses demonstrate that there are strong inflammatory effects resulting from FH inhibition. Notably, acute FH inhibition suppresses interleukin-10 expression, which leads to increased tumour necrosis factor secretion, an effect recapitulated by fumarate esters. Moreover, FH inhibition, but not fumarate esters, increases interferon-β production through mechanisms that are driven by mitochondrial RNA (mtRNA) release and activation of the RNA sensors TLR7, RIG-I and MDA5. This effect is recapitulated endogenously when FH is suppressed following prolonged lipopolysaccharide stimulation. Furthermore, cells from patients with systemic lupus erythematosus also exhibit FH suppression, which indicates a potential pathogenic role for this process in human disease. We therefore identify a protective role for FH in maintaining appropriate macrophage cytokine and interferon responses.
© 2023. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
Figures











Comment in
-
Mitochondrial fumarate implicated in inflammation.Nat Rev Rheumatol. 2023 May;19(5):257. doi: 10.1038/s41584-023-00963-z. Nat Rev Rheumatol. 2023. PMID: 37024712 No abstract available.
-
Fumarate hydratase: a new checkpoint of metabolic regulation in inflammatory macrophages.Signal Transduct Target Ther. 2023 Sep 4;8(1):332. doi: 10.1038/s41392-023-01594-2. Signal Transduct Target Ther. 2023. PMID: 37666844 Free PMC article. No abstract available.
References
-
- Jha AK et al. Network integration of parallel metabolic and transcriptional data reveals metabolic modules that regulate macrophage polarization. Immunity 42, 419–430 (2015). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

