Patient-derived spheroids and patient-derived organoids simulate evolutions of lung cancer
- PMID: 36895411
- PMCID: PMC9988482
- DOI: 10.1016/j.heliyon.2023.e13829
Patient-derived spheroids and patient-derived organoids simulate evolutions of lung cancer
Abstract
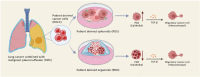
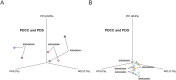
Cancer cells harbor many genetic mutations and gene expression profiles different from normal cells. Patient-derived cancer cells (PDCC) are preferred materials in cancer study. We established patient-derived spheroids (PDSs) and patient-derived organoids (PDOs) from PDCCs isolated from the malignant pleural effusion in 8 patients. The morphologies suggested that PDSs may be a model of local cancer extensions, while PDOs may be a model of distant cancer metastases. The gene expression profiles differed between PDSs and PDOs: Gene sets related to inflammatory responses and EMT were antithetically regulated in PDSs or in PDOs. PDSs demonstrated an attenuation of the pathways that contribute to the enhancement of transforming growth factor beta (TGF-β) induced epithelial mesenchymal transition (EMT), while PDOs demonstrated an attenuation of it. Taken together, PDSs and PDOs have differences in both the interaction to the immune systems and to the stroma. PDSs and PDOs will provide a model system that enable intimate investigation of the behavior of cancer cells in the body.
Keywords: Extracellular matrix; Immune system; Lung cancer; Organoid; Spheroid.
© 2023 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





References
-
- Amini L., Silbert S.K., Maude S.L., Nastoupil L.J., Ramos C.A., Brentjens R.J., Sauter C.S., Shah N.N., Abou-El-Enein M. Preparing for CAR T cell therapy: patient selection, bridging therapies and lymphodepletion. Nat. Rev. Clin. Oncol. 2022;19:342–355. - PubMed
-
- Basil M.C., Cardenas-Diaz F.L., Kathiriya J.J., Morley M.P., Carl J., Brumwell A.N., Katzen J., Slovik K.J., Babu A., Zhou S., Kremp M.M., McCauley K.B., Li S., Planer J.D., Hussain S.S., Liu X., Windmueller R., Ying Y., Stewart K.M., Oyster M., Christie J.D., Diamond J.M., Engelhardt J.F., Cantu E., Rowe S.M., Kotton D.N., Chapman H.A., Morrisey E.E. Human distal airways contain a multipotent secretory cell that can regenerate alveoli. Nature. 2022;604:120–126. - PMC - PubMed
LinkOut - more resources
Full Text Sources

