An optimal regulation of fluxes dictates microbial growth in and out of steady state
- PMID: 36896805
- PMCID: PMC10110240
- DOI: 10.7554/eLife.84878
An optimal regulation of fluxes dictates microbial growth in and out of steady state
Abstract
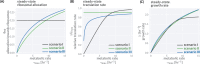
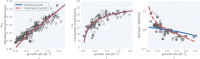
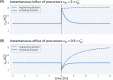
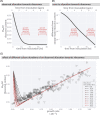
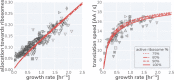
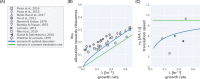
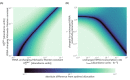
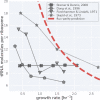
Effective coordination of cellular processes is critical to ensure the competitive growth of microbial organisms. Pivotal to this coordination is the appropriate partitioning of cellular resources between protein synthesis via translation and the metabolism needed to sustain it. Here, we extend a low-dimensional allocation model to describe the dynamic regulation of this resource partitioning. At the core of this regulation is the optimal coordination of metabolic and translational fluxes, mechanistically achieved via the perception of charged- and uncharged-tRNA turnover. An extensive comparison with ≈ 60 data sets from Escherichia coli establishes this regulatory mechanism's biological veracity and demonstrates that a remarkably wide range of growth phenomena in and out of steady state can be predicted with quantitative accuracy. This predictive power, achieved with only a few biological parameters, cements the preeminent importance of optimal flux regulation across conditions and establishes low-dimensional allocation models as an ideal physiological framework to interrogate the dynamics of growth, competition, and adaptation in complex and ever-changing environments.
Keywords: E. coli; S. cerevisiae; cell biology; infectious disease; microbial physiology; microbiology; physics of living systems; physiological modeling; resource allocation; systems biology.
© 2023, Chure and Cremer.
Conflict of interest statement
GC, JC No competing interests declared
Figures











Update of
- doi: 10.1101/2022.01.27.477569
Similar articles
-
Trade-offs between the instantaneous growth rate and long-term fitness: Consequences for microbial physiology and predictive computational models.Bioessays. 2023 Oct;45(10):e2300015. doi: 10.1002/bies.202300015. Epub 2023 Aug 9. Bioessays. 2023. PMID: 37559168 Review.
-
Cross-regulation between proteome reallocation and metabolic flux redistribution governs bacterial growth transition kinetics.Metab Eng. 2024 Mar;82:60-68. doi: 10.1016/j.ymben.2024.01.008. Epub 2024 Feb 2. Metab Eng. 2024. PMID: 38309620
-
A hybrid model of anaerobic E. coli GJT001: combination of elementary flux modes and cybernetic variables.Biotechnol Prog. 2008 Sep-Oct;24(5):993-1006. doi: 10.1002/btpr.73. Biotechnol Prog. 2008. PMID: 19194908
-
Metabolic-flux dependent regulation of microbial physiology.Curr Opin Microbiol. 2018 Apr;42:71-78. doi: 10.1016/j.mib.2017.10.029. Epub 2017 Nov 15. Curr Opin Microbiol. 2018. PMID: 29154077 Review.
-
Dynamical Allocation of Cellular Resources as an Optimal Control Problem: Novel Insights into Microbial Growth Strategies.PLoS Comput Biol. 2016 Mar 9;12(3):e1004802. doi: 10.1371/journal.pcbi.1004802. eCollection 2016 Mar. PLoS Comput Biol. 2016. PMID: 26958858 Free PMC article.
Cited by
-
Model-guided gene circuit design for engineering genetically stable cell populations in diverse applications.J R Soc Interface. 2025 Feb;22(223):20240602. doi: 10.1098/rsif.2024.0602. Epub 2025 Feb 12. J R Soc Interface. 2025. PMID: 39933591 Free PMC article.
-
Genome dilution by cell growth drives starvation-like proteome remodeling in mammalian and yeast cells.Nat Struct Mol Biol. 2024 Dec;31(12):1859-1871. doi: 10.1038/s41594-024-01353-z. Epub 2024 Jul 24. Nat Struct Mol Biol. 2024. PMID: 39048803
-
Functional regimes define soil microbiome response to environmental change.Nature. 2025 Aug;644(8078):1028-1038. doi: 10.1038/s41586-025-09264-9. Epub 2025 Jul 16. Nature. 2025. PMID: 40670792 Free PMC article.
-
Incoherent feedback from coupled amino acids and ribosome pools generates damped oscillations in growing E. coli.Nat Commun. 2025 Mar 29;16(1):3063. doi: 10.1038/s41467-025-57789-4. Nat Commun. 2025. PMID: 40157904 Free PMC article.
-
Systematic modulation of bacterial resource allocation by perturbing RNA polymerase availability via synthetic transcriptional switches.Nucleic Acids Res. 2025 Aug 11;53(15):gkaf814. doi: 10.1093/nar/gkaf814. Nucleic Acids Res. 2025. PMID: 40842238 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

