Structural Diversity among Edwardsiellaceae Core Oligosaccharides
- PMID: 36902212
- PMCID: PMC10003444
- DOI: 10.3390/ijms24054768
Structural Diversity among Edwardsiellaceae Core Oligosaccharides
Abstract
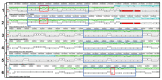
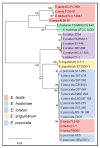
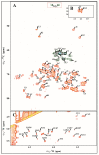
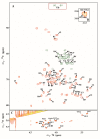
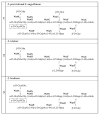
The Edwardsiella genus presents five different pathogenic species: Edwardsiella tarda, E. anguillarum, E. piscicida, E. hoshinae and E. ictaluri. These species cause infections mainly in fish, but they can also infect reptiles, birds or humans. Lipopolysaccharide (endotoxin) plays an important role in the pathogenesis of these bacteria. For the first time, the chemical structure and genomics of the lipopolysaccharide (LPS) core oligosaccharides of E. piscicida, E. anguillarum, E. hoshinae and E. ictaluri were studied. The complete gene assignments for all core biosynthesis gene functions were acquired. The structure of core oligosaccharides was investigated by ¹H and 13C nuclear magnetic resonance (NMR) spectroscopy. The structures of E. piscicida and E. anguillarum core oligosaccharides show the presence of →3,4)-L-glycero-α-D-manno-Hepp, two terminal β-D-Glcp, →2,3,7)-L-glycero-α-D-manno-Hepp, →7)-L-glycero-α-D-manno-Hepp, terminal α-D-GlcpN, two →4)-α-D-GalpA, → 3)-α-D-GlcpNAc, terminal β-D-Galp and →5-substituted Kdo. E. hoshinare core oligosaccharide shows only one terminal β-D-Glcp, and instead of terminal β-D-Galp a terminal α-D-GlcpNAc. E. ictaluri core oligosaccharide shows only one terminal β-D-Glcp, one →4)-α-D-GalpA and do not have terminal α-D-GlcpN (see complementary figure).
Keywords: Edwardsiellaea; NMR spectroscopy; core oligosaccharide; genomic.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Abbott S.L., Janda J.M. The genus Edwardsiella. Prokaryotes. 2006;6:72–89.
-
- Adeolu M., Alnajar S., Naushad S., Gupta R. Genome-based phylogeny and taxonomy of the ‘Enterobacteriales’: Proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov. Int. J. Syst. Evol. Microbiol. 2016;66:5575–5599. - PubMed
-
- Buján N., Mohammed H., Balboa S., Romalde J.L., Toranzo A.E., Arias C.R., Magariños B. Genetic studies to re-affiliate Edwardsiella tarda fish isolates to Edwardsiella piscicida and Edwardsiella anguillarum species. Syst. Appl. Microbiol. 2018;41:30–37. doi: 10.1016/j.syapm.2017.09.004. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

