Probing local changes to α-helical structures with 2D IR spectroscopy and isotope labeling
- PMID: 36906800
- PMCID: PMC10147839
- DOI: 10.1016/j.bpj.2023.03.014
Probing local changes to α-helical structures with 2D IR spectroscopy and isotope labeling
Abstract
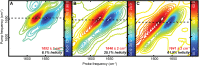
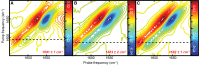
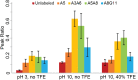
α-Helical secondary structures impart specific mechanical and physiochemical properties to peptides and proteins, enabling them to perform a vast array of molecular tasks ranging from membrane insertion to molecular allostery. Loss of α-helical content in specific regions can inhibit native protein function or induce new, potentially toxic, biological activities. Thus, identifying specific residues that exhibit loss or gain of helicity is critical for understanding the molecular basis of function. Two-dimensional infrared (2D IR) spectroscopy coupled with isotope labeling is capable of capturing detailed structural changes in polypeptides. Yet, questions remain regarding the inherent sensitivity of isotope-labeled modes to local changes in α-helicity, such as terminal fraying; the origin of spectral shifts (hydrogen-bonding versus vibrational coupling); and the ability to definitively detect coupled isotopic signals in the presence of overlapping side chains. Here, we address each of these points individually by characterizing a short, model α-helix (DPAEAAKAAAGR-NH2) with 2D IR and isotope labeling. These results demonstrate that pairs of 13C18O probes placed three residues apart can detect subtle structural changes and variations along the length of the model peptide as the α-helicity is systematically tuned. Comparison of singly and doubly labeled peptides affirm that frequency shifts arise primarily from hydrogen-bonding, while vibrational coupling between paired isotopes leads to increased peak areas that can be clearly differentiated from underlying side-chain modes or uncoupled isotope labels not participating in helical structures. These results demonstrate that 2D IR in tandem with i,i+3 isotope-labeling schemes can capture residue-specific molecular interactions within a single turn of an α-helix.
Copyright © 2023 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures


References
-
- Kuriyan J., Konforti B., Wemmer D. Garland Science; 2013. The Molecules of Life.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

