This is a preprint.
A Conserved Requirement for RME-8/DNAJC13 in Neuronal Autolysosome Reformation
- PMID: 36909501
- PMCID: PMC10002637
- DOI: 10.1101/2023.02.27.530319
A Conserved Requirement for RME-8/DNAJC13 in Neuronal Autolysosome Reformation
Update in
-
A conserved requirement for RME-8/DNAJC13 in neuronal autophagic lysosome reformation.Autophagy. 2024 Apr;20(4):792-808. doi: 10.1080/15548627.2023.2269028. Epub 2023 Nov 9. Autophagy. 2024. PMID: 37942902 Free PMC article.
Abstract
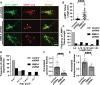
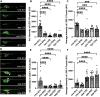
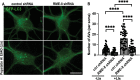
Autophagosomes fuse with lysosomes, forming autolysosomes that degrade engulfed cargo. To maintain lysosomal capacity, autolysosome reformation (ALR) must regenerate lysosomes from autolysosomes using a membrane tubule-based process. Maintaining lysosomal capacity is required to maintain proteostasis and cellular health, especially in neurons where lysosomal dysfunction has been repeatedly implicated in neurodegenerative disease. Cell biological studies have linked the DNA-J domain Hsc70 co-chaperone RME-8/DNAJC13 to endosomal coat protein regulation, while human genetics studies have linked RME-8/DNAJC13 to neurological disease, including Parkinsonism and Essential Tremor. We report new analysis of the requirements for the RME-8/DNAJC13 protein in neurons, focusing on C. elegans mechanosensory neurons in the intact animal, and in primary mouse cortical neurons in culture. We find that loss of RME-8/DNAJC13 in both systems results in accumulation of grossly elongated autolysosomal tubules. Further C. elegans analysis revealed a similar autolysosome tubule accumulation defect in mutants known to be required for ALR in mammals, including bec-1/beclin and vps-15/PIK3R4/p150 that regulate type-III PI3-kinase VPS-34, and dyn-1/dynamin that severs ALR tubules. Clathrin is also an important ALR regulator implicated in autolysosome tubule formation and release. In C. elegans we found that loss of RME-8 causes severe depletion of clathrin from neuronal autolysosomes, a phenotype shared with bec-1 and vps-15 mutants. We conclude that RME-8/DNAJC13 plays a conserved but previously unrecognized role in autolysosome reformation, likely affecting ALR tubule initiation and/or severing. Additionally, in both systems, we found that loss of RME-8/DNAJC13 appeared to reduce autophagic flux, suggesting feedback regulation from ALR to autophagy. Our results connecting RME-8/DNAJC13 to ALR and autophagy provide a potential mechanism by which RME-8/DNAJC13 could influence neuronal health and the progression of neurodegenerative disease.
Figures





References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
