Fully human monoclonal antibody targeting activated ADAM10 on colorectal cancer cells
- PMID: 36917886
- PMCID: PMC10499537
- DOI: 10.1016/j.biopha.2023.114494
Fully human monoclonal antibody targeting activated ADAM10 on colorectal cancer cells
Abstract
Metastasis and chemoresistance in colorectal cancer are mediated by certain poorly differentiated cancer cells, known as cancer stem cells, that are maintained by Notch downstream signaling initiated upon Notch cleavage by the metalloprotease ADAM10. It has been shown that ADAM10 overexpression correlates with aberrant signaling from Notch, erbBs, and other receptors, as well as a more aggressive metastatic phenotype, in a range of cancers including colon, gastric, prostate, breast, ovarian, uterine, and leukemia. ADAM10 inhibition, therefore, stands out as an important and new approach to deter the progression of advanced CRC. For targeting the ADAM10 substrate-binding region, which is located outside of the catalytic domain of the protease, we generated a human anti-ADAM10 monoclonal antibody named 1H5. Structural and functional characterization of 1H5 reveals that it binds to the substrate-binding cysteine-rich domain and recognizes an activated ADAM10 conformation present on tumor cells. The mAb inhibits Notch cleavage and proliferation of colon cancer cell lines in vitro and in mouse models. Consistent with its binding to activated ADAM10, the mAb augments the catalytic activity of ADAM10 towards small peptide substrates in vitro. Most importantly, in a mouse model of colon cancer, when administered in combination with the therapeutic agent Irinotecan, 1H5 causes highly effective tumor growth inhibition without any discernible toxicity effects. Our singular approach to target the ADAM10 substrate-binding region with therapeutic antibodies could overcome the shortcomings of previous intervention strategies of targeting the protease active site with small molecule inhibitors that exhibit musculoskeletal toxicity.
Keywords: ADAM10; COLO205; Chemotherapy; Colorectal cancer; Monoclonal antibody; Notch; Xenograft.
Copyright © 2023 The Authors. Published by Elsevier Masson SAS.. All rights reserved.
Conflict of interest statement
Conflict of Interest Statement The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


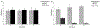
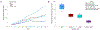

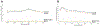


References
-
- Munro MJ, Wickremesekera SK, Peng L, Tan ST, Itinteang T, Cancer stem cells in colorectal cancer: a review, J. Clin. Pathol 71 (2) (2018) 110–116. - PubMed
-
- Hartmann DB, De Strooper B, Serneels L, Craessaerts K, Herreman A, Annaert W, Umans L, Lubke T, Lena Illert A, von Figura K, Saftig P, The disintegrin/metalloprotease ADAM 10 is essential for Notch signalling but not for alpha-secretase activity in fibroblasts, Hum. Mol. Genet 11 (2002) 2615–2624. - PubMed
-
- Murphy G, The ADAMs: signalling scissors in the tumour microenvironment, Nat. Rev. Cancer 8 (2008) 929–941. - PubMed
-
- Fischer M, Yen WC, Kapoun AM, Wang M, O’Young G, et al., Anti-DLL4 inhibits growth and reduces tumor-initiating cell frequency in colorectal tumors with oncogenic KRAS mutations, Cancer Res. 71 (2011) 1520–1525. - PubMed
-
- Li JL, Swanson RC, Oon CE, Turley H, Leek R, Sheldon H, Bridges E, et al., DLL4-Notch signaling mediates tumor resistance to anti-VEGF therapy in vivo, Cancer Res. 71 (2011) 6073–6083. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

