Depleting the 19S proteasome regulatory PSMD1 subunit as a cancer therapy strategy
- PMID: 36934426
- PMCID: PMC10225209
- DOI: 10.1002/cam4.5775
Depleting the 19S proteasome regulatory PSMD1 subunit as a cancer therapy strategy
Abstract
Background: Proteasome inhibitors are in use in treating certain types of cancers. These drugs inhibit the catalytic activity of the 20S proteasome, shared by all the different proteasome complexes. Inhibitors of the 26S-associated deubiquitinating activity explicitly inhibit the 26S proteasomal degradation of ubiquitinylated substrates. We have previously reported an alternative strategy that is based on reducing the 26S/20S ratio by depleting PSMD1, 6, and 11, the subunits of the 19S proteasome regulatory complex. Given the addiction of the many cancer types to a high 26S/20S ratio, the depletion strategy is highly effective in killing many aggressive cancer cell lines but not mouse and human immortalized and normal cells.
Methods: We used two aggressive cell lines, MDA-MB-231, a triple-negative breast tumor cell line, and OVCAR8, a high-grade ovary adenocarcinoma. Cell culture, mouse MDA-MB-231, OVCAR8 xenografts, and patient-derived ovarian cancer xenograft (PDX) models were transduced with lentivectors expressing PSMD1 shRNA. Tumor size was measured to follow treatment efficacy.
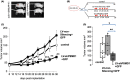
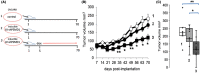
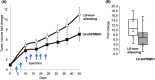
Results: Using different experimental strategies of expressing shRNA, we found that PSMD1 depletion, either by expressing PSMD1 shRNA in an inducible manner or in a constitutive manner, robustly inhibited MDA-MB-231, and OVCAR8 xenograft tumor growth. Furthermore, the PSMD1 depletion strategy compromised the growth of the PDX of primary ovarian cancer.
Conclusion: Our results suggest that reducing the 26S/20S ratio might be a valuable strategy for treating drug-resistant aggressive types of cancers.
Keywords: 19S regulatory particle; 26S proteasome; breast cancer; cancer therapy; mouse xenografts; ovarian cancer PDX.
© 2023 The Authors. Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Oncogenic addiction to high 26S proteasome level.Cell Death Dis. 2018 Jul 10;9(7):773. doi: 10.1038/s41419-018-0806-4. Cell Death Dis. 2018. PMID: 29991718 Free PMC article.
-
Method of Monitoring 26S Proteasome in Cells Revealed the Crucial Role of PSMA3 C-Terminus in 26S Integrity.Biomolecules. 2023 Jun 15;13(6):992. doi: 10.3390/biom13060992. Biomolecules. 2023. PMID: 37371572 Free PMC article.
-
Depletion of proteasome subunit PSMD1 induces cancer cell death via protein ubiquitination and DNA damage, irrespective of p53 status.Sci Rep. 2024 Apr 5;14(1):7997. doi: 10.1038/s41598-024-58215-3. Sci Rep. 2024. PMID: 38580756 Free PMC article.
-
Redox regulation of the proteasome via S-glutathionylation.Redox Biol. 2013 Dec 14;2:44-51. doi: 10.1016/j.redox.2013.12.003. Redox Biol. 2013. PMID: 24396728 Free PMC article. Review.
-
Proteasomes of the yeast S. cerevisiae: genes, structure and functions.Mol Biol Rep. 1995;21(1):3-10. doi: 10.1007/BF00990964. Mol Biol Rep. 1995. PMID: 7565661 Review.
Cited by
-
[Research and Therapeutic Advances of 26S Proteasome Subunit in Non-small Cell Lung Cancer].Zhongguo Fei Ai Za Zhi. 2025 May 20;28(5):363-370. doi: 10.3779/j.issn.1009-3419.2025.106.13. Zhongguo Fei Ai Za Zhi. 2025. PMID: 40506490 Free PMC article. Review. Chinese.
-
Molecular subtype identification and prognosis stratification based on lysosome-related genes in breast cancer.Heliyon. 2024 Feb 7;10(4):e25643. doi: 10.1016/j.heliyon.2024.e25643. eCollection 2024 Feb 29. Heliyon. 2024. PMID: 38420434 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

