Effects of allicin on human Simpson-Golabi-Behmel syndrome cells in mediating browning phenotype
- PMID: 36936145
- PMCID: PMC10014806
- DOI: 10.3389/fendo.2023.1141303
Effects of allicin on human Simpson-Golabi-Behmel syndrome cells in mediating browning phenotype
Abstract
Introduction: Obesity is a major health problem because it is associated with increased risk of cardiovascular disease, diabetes, hypertension, and some cancers. Strategies to prevent or reduce obesity focus mainly on the possible effects of natural compounds that can induce a phenotype of browning adipocytes capable of releasing energy in the form of heat. Allicin, a bioactive component of garlic with numerous pharmacological functions, is known to stimulate energy metabolism.
Methods: In the present study, the effects of allicin on human Simpson-Golabi-Behmel Syndrome (SGBS) cells were investigated by quantifying the dynamics of lipid droplets (LDs) and mitochondria, as well as transcriptomic changes after six days of differentiation.
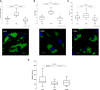
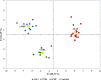
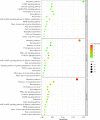
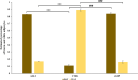
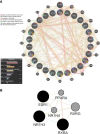
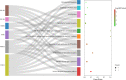
Results: Allicin significantly promoted the reduction in the surface area and size of LDs, leading to the formation of multilocular adipocytes, which was confirmed by the upregulation of genes related to lipolysis. The increase in the number and decrease in the mean aspect ratio of mitochondria in allicin-treated cells indicate a shift in mitochondrial dynamics toward fission. The structural results are confirmed by transcriptomic analysis showing a significant arrangement of gene expression associated with beige adipocytes, in particular increased expression of T-box transcription factor 1 (TBX1), uncoupling protein 1 (UCP1), PPARG coactivator 1 alpha (PPARGC1A), peroxisome proliferator-activated receptor alpha (PPARA), and OXPHOS-related genes. The most promising targets are nuclear genes such as retinoid X receptor alpha (RXRA), retinoid X receptor gamma (RXRG), nuclear receptor subfamily 1 group H member 3 (NR1H3), nuclear receptor subfamily 1 group H member 4 (NR1H4), PPARA, and oestrogen receptor 1 (ESR1).
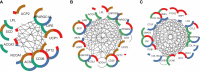
Discussion: Transcriptomic data and the network pharmacology-based approach revealed that genes and potential targets of allicin are involved in ligand-activated transcription factor activity, intracellular receptor signalling, regulation of cold-induced thermogenesis, and positive regulation of lipid metabolism. The present study highlights the potential role of allicin in triggering browning in human SGBS cells by affecting the LD dynamics, mitochondrial morphology, and expression of brown marker genes. Understanding the potential targets through which allicin promotes this effect may reveal the underlying signalling pathways and support these findings.
Keywords: SGBS cells; allicin; lipid droplets; mitochondria; thermogenesis.
Copyright © 2023 Ali, Wabitsch, Tews and Colitti.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The handling editor EKK declared a past co-authorship with the author MW.
Figures



Similar articles
-
Differentiating SGBS adipocytes respond to PPARγ stimulation, irisin and BMP7 by functional browning and beige characteristics.Sci Rep. 2019 Apr 9;9(1):5823. doi: 10.1038/s41598-019-42256-0. Sci Rep. 2019. PMID: 30967578 Free PMC article.
-
Simpson-Golabi-Behmel syndrome human adipocytes reveal a changing phenotype throughout differentiation.Histochem Cell Biol. 2018 Jun;149(6):593-605. doi: 10.1007/s00418-018-1663-z. Epub 2018 Mar 24. Histochem Cell Biol. 2018. PMID: 29574488
-
Clozapine modifies the differentiation program of human adipocytes inducing browning.Transl Psychiatry. 2016 Nov 29;6(11):e963. doi: 10.1038/tp.2016.230. Transl Psychiatry. 2016. PMID: 27898069 Free PMC article.
-
The ménage à trois of autophagy, lipid droplets and liver disease.Autophagy. 2022 Jan;18(1):50-72. doi: 10.1080/15548627.2021.1895658. Epub 2021 Apr 2. Autophagy. 2022. PMID: 33794741 Free PMC article. Review.
-
Positive and negative control of Ucp1 gene transcription and the role of β-adrenergic signaling networks.Int J Obes (Lond). 2010 Oct;34 Suppl 1:S28-33. doi: 10.1038/ijo.2010.180. Int J Obes (Lond). 2010. PMID: 20935662 Review.
Cited by
-
Anti-obesity peptides from food: Production, evaluation, sources, and commercialization.Compr Rev Food Sci Food Saf. 2025 Mar;24(2):e70158. doi: 10.1111/1541-4337.70158. Compr Rev Food Sci Food Saf. 2025. PMID: 40111015 Free PMC article. Review.
-
Complex networks interactions between bioactive compounds and adipose tissue vis-à-vis insulin resistance.Front Endocrinol (Lausanne). 2025 May 13;16:1578552. doi: 10.3389/fendo.2025.1578552. eCollection 2025. Front Endocrinol (Lausanne). 2025. PMID: 40433407 Free PMC article.
References
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

