A c-di-GMP binding effector controls cell size in a cyanobacterium
- PMID: 36947515
- PMCID: PMC10068817
- DOI: 10.1073/pnas.2221874120
A c-di-GMP binding effector controls cell size in a cyanobacterium
Abstract
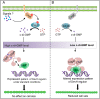
Cyclic-di-GMP (c-di-GMP) is a ubiquitous bacterial signaling molecule. It is also a critical player in the regulation of cell size and cell behaviors such as cell aggregation and phototaxis in cyanobacteria, which constitute an important group of prokaryotes for their roles in the ecology and evolution of the Earth. However, c-di-GMP receptors have never been revealed in cyanobacteria. Here, we report the identification of a c-di-GMP receptor, CdgR, from the filamentous cyanobacterium Anabaena PCC 7120. Crystal structural analysis and genetic studies demonstrate that CdgR binds c-di-GMP at the dimer interface and this binding is required for the control of cell size in a c-di-GMP-dependent manner. Different functions of CdgR, in ligand binding and signal transmission, could be separated genetically, allowing us to dissect its molecular signaling functions. The presence of the apo-form of CdgR triggers cell size reduction, consistent with the similar effects observed with a decrease of c-di-GMP levels in cells. Furthermore, we found that CdgR exerts its function by interacting with a global transcription factor DevH, and this interaction was inhibited by c-di-GMP. The lethal effect triggered by conditional depletion of DevH or by the production of several point-mutant proteins of CdgR in cells indicates that this signaling pathway plays critical functions in Anabaena. Our studies revealed a mechanism of c-di-GMP signaling in the control of cell size, an important and complex trait for bacteria. CdgR is highly conserved in cyanobacteria, which will greatly expand our understanding of the roles of c-di-GMP signaling in these organisms.
Keywords: c-di-GMP; c-di-GMP effector; cell size; cyanobacteria; signal transduction.
Conflict of interest statement
The authors declare no competing interest.
Figures




Similar articles
-
Control of Cell Size by c-di-GMP Requires a Two-Component Signaling System in the Cyanobacterium Anabaena sp. Strain PCC 7120.Microbiol Spectr. 2023 Feb 14;11(1):e0422822. doi: 10.1128/spectrum.04228-22. Epub 2023 Jan 10. Microbiol Spectr. 2023. PMID: 36625639 Free PMC article.
-
Occurrence of cyclic di-GMP-modulating output domains in cyanobacteria: an illuminating perspective.mBio. 2013 Aug 13;4(4):e00451-13. doi: 10.1128/mBio.00451-13. mBio. 2013. PMID: 23943760 Free PMC article.
-
Cyanobacteriochrome SesA is a diguanylate cyclase that induces cell aggregation in Thermosynechococcus.J Biol Chem. 2014 Sep 5;289(36):24801-9. doi: 10.1074/jbc.M114.583674. Epub 2014 Jul 24. J Biol Chem. 2014. PMID: 25059661 Free PMC article.
-
Diversity of Cyclic Di-GMP-Binding Proteins and Mechanisms.J Bacteriol. 2016 Jan 1;198(1):32-46. doi: 10.1128/JB.00333-15. J Bacteriol. 2016. PMID: 26055114 Free PMC article. Review.
-
Biofilm control by interfering with c-di-GMP metabolism and signaling.Biotechnol Adv. 2022 May-Jun;56:107915. doi: 10.1016/j.biotechadv.2022.107915. Epub 2022 Jan 31. Biotechnol Adv. 2022. PMID: 35101567 Review.
Cited by
-
ComFB, a new widespread family of c-di-NMP receptor proteins.bioRxiv [Preprint]. 2024 Nov 10:2024.11.10.622515. doi: 10.1101/2024.11.10.622515. bioRxiv. 2024. PMID: 39574629 Free PMC article. Preprint.
-
Gas and light: triggers of c-di-GMP-mediated regulation.FEMS Microbiol Rev. 2023 Jul 5;47(4):fuad034. doi: 10.1093/femsre/fuad034. FEMS Microbiol Rev. 2023. PMID: 37339911 Free PMC article. Review.
-
Control of light-dependent behaviour in cyanobacteria by the second messenger cyclic di-GMP.Microlife. 2023 Apr 12;4:uqad019. doi: 10.1093/femsml/uqad019. eCollection 2023. Microlife. 2023. PMID: 37223735 Free PMC article. Review.
-
A novel transcriptional regulator, CdeR, modulates the type III secretion system via c-di-GMP signaling in Dickeya dadantii.Microbiol Spectr. 2025 Apr;13(4):e0265524. doi: 10.1128/spectrum.02655-24. Epub 2025 Mar 5. Microbiol Spectr. 2025. PMID: 40042333 Free PMC article.
-
Metabolic activities of marine ammonia-oxidizing archaea orchestrated by quorum sensing.mLife. 2024 Sep 30;3(3):417-429. doi: 10.1002/mlf2.12144. eCollection 2024 Sep. mLife. 2024. PMID: 39359677 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

