Conformation of influenza AM2 membrane protein in nanodiscs and liposomes
- PMID: 36948480
- PMCID: PMC10175228
- DOI: 10.1016/j.bbamem.2023.184152
Conformation of influenza AM2 membrane protein in nanodiscs and liposomes
Abstract
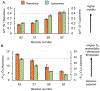
The influenza A M2 protein (AM2) is a multifunctional membrane-associated homotetramer that orchestrates several essential events in the viral infection cycle including viral assembly and budding. An atomic-level conformational understanding of this key player in the influenza life cycle could inform new antiviral strategies. For conformational studies of complex systems like the AM2 membrane protein, a multipronged approach using different biophysical methods and different model membranes is a powerful way to incorporate complementary data and achieve a fuller, more robust understanding of the system. However, one must be aware of how the sample composition required for a particular method impacts the data collected and how conclusions are drawn. In that spirit, we systematically compared the properties of AM2 in two different model membranes: nanodiscs and liposomes. Electron paramagnetic spectroscopy of spin-labeled AM2 showed that the conformation and dynamics were strikingly similar in both AM2-nanodiscs and AM2-liposomes consistent with similar conformations in both model membranes. Analysis of spin labeled lipids embedded in both model membranes revealed that the bilayer in AM2-liposomes was more fluid and permeable to oxygen than AM2-nanodiscs with the same lipid composition. Once the difference in the partitioning of the paramagnetic oxygen relaxation agent was taken into account, the membrane topology of AM2 appeared to be the same in both liposomes and nanodiscs. Finally, functionally relevant AM2 conformational shifts previously seen in liposomes due to the addition of cholesterol were also observed in nanodiscs.
Keywords: Electron paramagnetic resonance; Influenza A M2 protein; Liposomes; Membrane proteins; Nanodiscs; Oxygen concentration in membranes; Site-directed spin labeling.
Copyright © 2023 Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Kathleen P. Howard reports financial support was provided by National Institutes of Health.
Figures


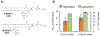
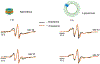
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

