Super-enhancer-associated transcription factors collaboratively regulate trophoblast-active gene expression programs in human trophoblast stem cells
- PMID: 36951126
- PMCID: PMC10164554
- DOI: 10.1093/nar/gkad215
Super-enhancer-associated transcription factors collaboratively regulate trophoblast-active gene expression programs in human trophoblast stem cells
Abstract
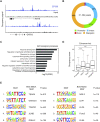
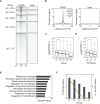
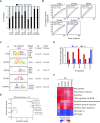
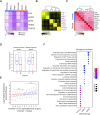
The placenta is an essential organ that supports the growth and development of the fetus during pregnancy. However, cell type-specific enhancers and transcription factors (TFs), and the mechanisms underlying the maintenance and differentiation of trophoblast stem cell (TSC) populations in the human placenta remain elusive. Here, using human TSCs as a model system, we identify 31,362 enhancers that are enriched with the motifs of previously reported TSC-pivotal TFs, including TEAD4, GATA2/3 and TFAP2C. Subsequently, we identify 580 super-enhancers (SEs) and 549 SE-associated genes. These genes are robustly expressed in the human placenta and include numerous TFs, implying that SE-associated TFs (SE-TFs) may play crucial roles in placental development. Additionally, we identify the global binding sites of five TSC-pivotal SE-TFs (FOS, GATA2, MAFK, TEAD4 and TFAP2C), revealing that they preferentially co-occupy enhancers, regulate each other and form a trophoblast-active gene regulatory network. Loss-of-function studies unveil that the five TFs promote self-renewal of TSCs by activating proliferation-associated genes while repressing developmental genes. We further reveal that the five TFs exert conserved and unique functions on placental development between humans and mice. Our study provides important insights into the roles of human TSC-pivotal TFs in regulating placenta-specific gene expression programs.
© The Author(s) 2023. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


References
-
- Maltepe E., Fisher S.J.. Placenta: the forgotten organ. Annu. Rev. Cell Dev. Biol. 2015; 31:523–552. - PubMed
-
- Turco M.Y., Moffett A.. Development of the human placenta. Development. 2019; 146:dev163428. - PubMed
-
- Okae H., Toh H., Sato T., Hiura H., Takahashi S., Shirane K., Kabayama Y., Suyama M., Sasaki H., Arima T.. Derivation of human trophoblast stem cells. Cell Stem Cell. 2018; 22:50–63. - PubMed
-
- Lemon B., Tjian R.. Orchestrated response: a symphony of transcription factors for gene control. Genes Dev. 2000; 14:2551–2569. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

