Structural Basis for Binding of Neutralizing Antibodies to Clostridioides difficile Binary Toxin
- PMID: 36951574
- PMCID: PMC10127577
- DOI: 10.1128/jb.00456-22
Structural Basis for Binding of Neutralizing Antibodies to Clostridioides difficile Binary Toxin
Abstract
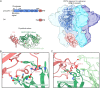
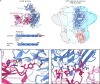
Clostridioides difficile is a Gram-positive opportunistic human pathogen that causes 15,000 deaths annually in the United States, prompting a need for vaccine development. In addition to the important toxins TcdA and TcdB, binary toxin (CDT) plays a significant role in the pathogenesis of certain C. difficile ribotypes by catalyzing the ADP-ribosylation of actin in host cells. However, the mechanisms of CDT neutralization by antibodies have not been studied, limiting our understanding of key epitopes for CDT antigen design. Therefore, we isolated neutralizing monoclonal antibodies against CDT and characterized their mechanisms of neutralization structurally and biochemically. Here, 2.5-Å and 2.6-Å resolution X-ray crystal structures of the antibodies BINTOXB/22 and BINTOXB/9, respectively, in complex with CDTb-the CDT subunit that forms a heptameric pore for the delivery of toxic CDTa enzyme into the host cytosol-showed that both antibodies sterically clash with adjacent protomers in the assembled heptamer. Assessment of trypsin-induced oligomerization of the purified CDTb protoxin in vitro showed that BINTOXB/22 and BINTOXB/9 prevented the assembly of di-heptamers upon prodomain cleavage. This work suggests that the CDT oligomerization process can be effectively targeted by antibodies, which will aid in the development of C. difficile vaccines and therapeutics. IMPORTANCE Clostridioides difficile strains associated with worse clinical outcomes have been found to secrete a toxin called CDT (or binary toxin). As blocking the function of this toxin could help mitigate C. difficile infections, we sought to determine the molecular basis for the inhibition of CDT by monoclonal antibodies. We isolated monoclonal antibodies targeting the B-component of CDT (CDTb) and selected two with neutralizing activity for detailed structural and biochemical characterization. High-resolution crystal structures of each antibody bound to CDTb showed that their presence would preclude the assembly of a CDTb oligomer required for activity. Oligomerization of CDTb in vitro was shown to be blocked in the presence of the neutralizing antibodies, but not a control antibody.
Keywords: Clostridioides difficile; X-ray crystallography; neutralizing antibodies; pore-forming toxins.
Conflict of interest statement
The authors declare a conflict of interest. N.B., P.H., and V.D. are current or former employees of the GlaxoSmithKline group of companies. N.B. is currently employed at Biovectra, a CDMO and API Manufacturing company.
Figures




Similar articles
-
The cytotoxic effect of Clostridioides difficile pore-forming toxin CDTb.Biochim Biophys Acta Biomembr. 2021 Jun 1;1863(6):183603. doi: 10.1016/j.bbamem.2021.183603. Epub 2021 Mar 6. Biochim Biophys Acta Biomembr. 2021. PMID: 33689753
-
Structure of the cell-binding component of the Clostridium difficile binary toxin reveals a di-heptamer macromolecular assembly.Proc Natl Acad Sci U S A. 2020 Jan 14;117(2):1049-1058. doi: 10.1073/pnas.1919490117. Epub 2020 Jan 2. Proc Natl Acad Sci U S A. 2020. PMID: 31896582 Free PMC article.
-
The Importance of Therapeutically Targeting the Binary Toxin from Clostridioides difficile.Int J Mol Sci. 2021 Mar 13;22(6):2926. doi: 10.3390/ijms22062926. Int J Mol Sci. 2021. PMID: 33805767 Free PMC article. Review.
-
Structural elucidation of the Clostridioides difficile transferase toxin reveals a single-site binding mode for the enzyme.Proc Natl Acad Sci U S A. 2020 Mar 17;117(11):6139-6144. doi: 10.1073/pnas.1920555117. Epub 2020 Mar 2. Proc Natl Acad Sci U S A. 2020. PMID: 32123082 Free PMC article.
-
Clostridium difficile binary toxin CDT: mechanism, epidemiology, and potential clinical importance.Gut Microbes. 2014 Jan-Feb;5(1):15-27. doi: 10.4161/gmic.26854. Epub 2013 Oct 31. Gut Microbes. 2014. PMID: 24253566 Free PMC article. Review.
Cited by
-
New treatment approaches for Clostridioides difficile infections: alternatives to antibiotics and fecal microbiota transplantation.Gut Microbes. 2024 Jan-Dec;16(1):2337312. doi: 10.1080/19490976.2024.2337312. Epub 2024 Apr 9. Gut Microbes. 2024. PMID: 38591915 Free PMC article. Review.
-
Structural and Functional Insights into the Delivery Systems of Bacillus and Clostridial Binary Toxins.Toxins (Basel). 2024 Jul 25;16(8):330. doi: 10.3390/toxins16080330. Toxins (Basel). 2024. PMID: 39195740 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

