Construction of copy number variation landscape and characterization of associated genes in a Bangladeshi cohort of neurodevelopmental disorders
- PMID: 36959829
- PMCID: PMC10028086
- DOI: 10.3389/fgene.2023.955631
Construction of copy number variation landscape and characterization of associated genes in a Bangladeshi cohort of neurodevelopmental disorders
Abstract
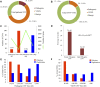
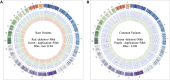
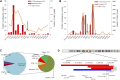
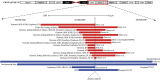
Introduction: Copy number variations (CNVs) play a critical role in the pathogenesis of neurodevelopmental disorders (NDD) among children. In this study, we aim to identify clinically relevant CNVs, genes and their phenotypic characteristics in an ethnically underrepresented homogenous population of Bangladesh. Methods: We have conducted chromosomal microarray analysis (CMA) for 212 NDD patients with male to female ratio of 2.2:1.0 to identify rare CNVs. To identify candidate genes within the rare CNVs, gene constraint metrics [i.e., "Critical-Exon Genes (CEGs)"] were applied to the population data. Autism Diagnostic Observation Schedule-Second Edition (ADOS-2) was followed in a subset of 95 NDD patients to assess the severity of autism and all statistical tests were performed using the R package. Results: Of all the samples assayed, 12.26% (26/212) and 57.08% (121/212) patients carried pathogenic and variant of uncertain significance (VOUS) CNVs, respectively. While 2.83% (6/212) patients' pathogenic CNVs were found to be located in the subtelomeric regions. Further burden test identified females are significant carriers of pathogenic CNVs compared to males (OR = 4.2; p = 0.0007). We have observed an increased number of Loss of heterozygosity (LOH) within cases with 23.85% (26/109) consanguineous parents. Our analyses on imprinting genes show, 36 LOH variants disrupting 69 unique imprinted genes and classified these variants as VOUS. ADOS-2 subset shows severe social communication deficit (p = 0.014) and overall ASD symptoms severity (p = 0.026) among the patients carrying duplication CNV compared to the CNV negative group. Candidate gene analysis identified 153 unique CEGs in pathogenic CNVs and 31 in VOUS. Of the unique genes, 18 genes were found to be in smaller (<1 MB) focal CNVs in our NDD cohort and we identified PSMC3 gene as a strong candidate gene for Autism Spectrum Disorder (ASD). Moreover, we hypothesized that KMT2B gene duplication might be associated with intellectual disability. Conclusion: Our results show the utility of CMA for precise genetic diagnosis and its integration into the diagnosis, therapy and management of NDD patients.
Keywords: Autism Diagnostic Observation Schedule-Second Edition (ADOS-2); autism spectrum disorder (ASD); chromosomal microarray analysis (CMA); copy number variation (CNV); critical exon gene (CEG); neurodevelopmental disorders (NDDs); variant of uncertain significance (VOUS).
Copyright © 2023 Akter, Rahman, Sarker, Basiruzzaman, Islam, Rahaman, Rahaman, Eshaque, Dity, Sarker, Amin, Hossain, Lopa, Jahan, Hossain, Islam, Mondol, Faruk, Saha, Kundu, Kanta, Kazal, Fatema, Rahman, Hasan, Hossain Mollah, Hosen, Karuvantevida, Begum, Zehra, Nassir, Nabi, Uddin and Uddin.
Conflict of interest statement
Authors AI and MU are employees of GenomeArc Inc. Authors HA, SLS, MB, MMI, MAR(1), MAR(2), TBE, NJD, ML, NJ, AI, AM, MOF, and KMFU are employees of NeuroGen Healthcare. The authors declare that this study received funding from NeuroGen Healthcare. The funder was not involved in the study design, collection, analysis, interpretation of data, and the writing of this article or the decision to submit it for publication. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Abou Jamra R., Wohlfart S., Zweier M., Uebe S., Priebe L., Ekici A., et al. (2011). Homozygosity mapping in 64 Syrian consanguineous families with non-specific intellectual disability reveals 11 novel loci and high heterogeneity. Eur. J. Hum. Genet. 19, 1161–1166. 10.1038/ejhg.2011.98 - DOI - PMC - PubMed
-
- Akter H., Hossain M. S., Dity N. J., Rahaman M. A., Furkan Uddin K. M., Nassir N., et al. (2021). Whole exome sequencing uncovered highly penetrant recessive mutations for a spectrum of rare genetic pediatric diseases in Bangladesh. npj Genomic Med. 6, 14–19. 10.1038/s41525-021-00173-0 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

