Developmental dynamic transcriptome and systematic analysis reveal the major genes underlying isoflavone accumulation in soybean
- PMID: 36959940
- PMCID: PMC10027745
- DOI: 10.3389/fpls.2023.1014349
Developmental dynamic transcriptome and systematic analysis reveal the major genes underlying isoflavone accumulation in soybean
Abstract
Introduction: Soy isoflavone, a class of polyphenolic compounds exclusively occurred in legumes, is an important bioactive compound for both plants and human beings. The outline of isoflavones biosynthesis pathway has been drawn up basically in the previous research. However, research on the subject has been mostly restricted to investigate the static regulation of isoflavone content in soybean, rather than characterize its dynamic variation and modulation network in developing seeds.
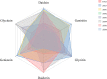
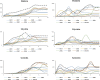
Methods: In this study, by using eight recombinant inbred lines (RIL), the contents of six isoflavone components in the different stages of developing soybean seeds were determined to characterize the dynamic variation of isoflavones, and the isoflavones accumulation pattern at physiological level was investigated. Meanwhile, we integrated and analyzed the whole genome expression profile of four lines and 42 meta-transcriptome data, based on the multiple algorithms.
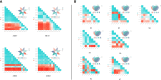
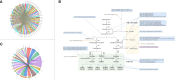
Results: This study: 1) obtained 4 molecular modules strongly correlated with isoflavone accumulation; 2) identified 28 novel major genes that could affect the accumulation of isoflavones in developing seeds free from the limitation of environments; 3) discussed the dynamic molecular patterns regulating isoflavones accumulation in developing seed; 4) expanded the isoflavone biosynthesis pathway.
Discussion: The results not only promote the understandings on the biosynthesis and regulation of isoflavones at physiological and molecular level, but also facilitate to breed elite soybean cultivars with high isoflavone contents.
Keywords: dynamic accumulation patterns; dynamic regulation networks; meta-analysis; soy isoflavone; transcriptome.
Copyright © 2023 Chen, Liu, Li, Wang, Pan, Wang and Zhang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
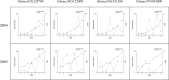
Figures

Similar articles
-
Contextualized Metabolic Modelling Revealed Factors Affecting Isoflavone Accumulation in Soybean Seeds.Plant Cell Environ. 2024 Sep 18. doi: 10.1111/pce.15140. Online ahead of print. Plant Cell Environ. 2024. PMID: 39292176
-
Genome wide association study to detect genetic regions related to isoflavone content in a mutant soybean population derived from radiation breeding.Front Plant Sci. 2022 Aug 18;13:968466. doi: 10.3389/fpls.2022.968466. eCollection 2022. Front Plant Sci. 2022. PMID: 36061785 Free PMC article.
-
MATE-Type Proteins Are Responsible for Isoflavone Transportation and Accumulation in Soybean Seeds.Int J Mol Sci. 2021 Nov 6;22(21):12017. doi: 10.3390/ijms222112017. Int J Mol Sci. 2021. PMID: 34769445 Free PMC article.
-
The association between soy-based food and soy isoflavone intake and the risk of gastric cancer: a systematic review and meta-analysis.J Sci Food Agric. 2021 Oct;101(13):5314-5324. doi: 10.1002/jsfa.11334. Epub 2021 Jun 16. J Sci Food Agric. 2021. PMID: 34032287
-
Occurrence of isoflavones in soybean sprouts and strategies to enhance their content: A review.J Food Sci. 2022 May;87(5):1961-1982. doi: 10.1111/1750-3841.16131. Epub 2022 Apr 11. J Food Sci. 2022. PMID: 35411587 Review.
Cited by
-
Identifying Candidate Genes Related to Soybean (Glycine max) Seed Coat Color via RNA-Seq and Coexpression Network Analysis.Genes (Basel). 2025 Jan 1;16(1):44. doi: 10.3390/genes16010044. Genes (Basel). 2025. PMID: 39858589 Free PMC article.
-
Bioinformatics Identification and Expression Analysis of Acetyl-CoA Carboxylase Reveal Its Role in Isoflavone Accumulation during Soybean Seed Development.Int J Mol Sci. 2024 Sep 23;25(18):10221. doi: 10.3390/ijms251810221. Int J Mol Sci. 2024. PMID: 39337707 Free PMC article.
References
-
- Borenstein M., Hedges L. V., Higgins J. P. T., Rothstein H. R. (2009). How a meta-analysis works. Introduction to Meta-Anal. 1–7. doi: 10.1002/9780470743386.ch1 - DOI
-
- Dhaubhadel S., McGarvey B. D., Williams R., Gijzen M. (2003). Isoflavonoid biosynthesis and accumulation in developing soybean seeds. Plant Mol. Biol. 53, 733–743. doi: 10.1023/B:PLAN.0000023666.30358.ae - DOI - PubMed
LinkOut - more resources
Full Text Sources

