Individualised human phenotype ontology gene panels improve clinical whole exome and genome sequencing analytical efficacy in a cohort of developmental and epileptic encephalopathies
- PMID: 36967109
- PMCID: PMC10337286
- DOI: 10.1002/mgg3.2167
Individualised human phenotype ontology gene panels improve clinical whole exome and genome sequencing analytical efficacy in a cohort of developmental and epileptic encephalopathies
Abstract
Background: The majority of genetic epilepsies remain unsolved in terms of specific genotype. Phenotype-based genomic analyses have shown potential to strengthen genomic analysis in various ways, including improving analytical efficacy.
Methods: We have tested a standardised phenotyping method termed 'Phenomodels' for integrating deep-phenotyping information with our in-house developed clinical whole exome/genome sequencing analytical pipeline. Phenomodels includes a user-friendly epilepsy phenotyping template and an objective measure for selecting which template terms to include in individualised Human Phenotype Ontology (HPO) gene panels. In a pilot study of 38 previously solved cases of developmental and epileptic encephalopathies, we compared the sensitivity and specificity of the individualised HPO gene panels with the clinical epilepsy gene panel.
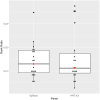
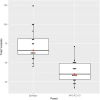
Results: The Phenomodels template showed high sensitivity for capturing relevant phenotypic information, where 37/38 individuals' HPO gene panels included the causative gene. The HPO gene panels also had far fewer variants to assess than the epilepsy gene panel.
Conclusion: We have demonstrated a viable approach for incorporating standardised phenotype information into clinical genomic analyses, which may enable more efficient analysis.
Keywords: deep phenotyping; epilepsy; human phenotype ontology; precision medicine; whole exome sequencing; whole genome sequencing.
© 2023 The Authors. Molecular Genetics & Genomic Medicine published by Wiley Periodicals LLC.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
-
- Akawi, N. , McRae, J. , Ansari, M. , Balasubramanian, M. , Blyth, M. , Brady, A. F. , Clayton, S. , Cole, T. , Deshpande, C. , Fitzgerald, T. W. , Foulds, N. , Francis, R. , Gabriel, G. , Gerety, S. S. , Goodship, J. , Hobson, E. , Jones, W. D. , Joss, S. , King, D. , … DDD Study . (2015). Discovery of four recessive developmental disorders using probabilistic genotype and phenotype matching among 4,125 families. Nature Genetics, 47(11), 1363–1369. 10.1038/ng.3410 - DOI - PMC - PubMed
-
- Allen, N. M. , Conroy, J. , Shahwan, A. , Lynch, B. , Correa, R. G. , Pena, S. D. , McCreary, D. , Magalhaes, T. R. , Ennis, S. , Lynch, S. A. , & King, M. D. (2016). Unexplained early onset epileptic encephalopathy: Exome screening and phenotype expansion. Epilepsia, 57(1), e12–e17. 10.1111/epi.13250 - DOI - PubMed
-
- Bastarache, L. , Hughey, J. J. , Hebbring, S. , Marlo, J. , Zhao, W. , Ho, W. T. , Van Driest, S. L. , McGregor, T. L. , Mosley, J. D. , Wells, Q. S. , Temple, M. , Ramirez, A. H. , Carroll, R. , Osterman, T. , Edwards, T. , Ruderfer, D. , Velez Edwards, D. R. , Hamid, R. , Cogan, J. , … Denny, J. C. (2018). Phenotype risk scores identify patients with unrecognized Mendelian disease patterns. Science, 359(6381), 1233–1239. 10.1126/science.aal4043 - DOI - PMC - PubMed
-
- Bennett, M. F. , Hildebrand, M. S. , Kayumi, S. , Corbett, M. A. , Gupta, S. , Ye, Z. , Krivanek, M. , Burgess, R. , Henry, O. J. , Damiano, J. A. , Boys, A. , Gecz, J. , Bahlo, M. , Scheffer, I. E. , & Berkovic, S. F. (2022). Evidence for a dual‐pathway, 2‐hit genetic model for focal cortical dysplasia and Epilepsy. Neurology Genetics, 8(1), e652. 10.1212/NXG.0000000000000652 - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

