Knocking Down CDKN2A in 3D hiPSC-Derived Brown Adipose Progenitors Potentiates Differentiation, Oxidative Metabolism and Browning Process
- PMID: 36980212
- PMCID: PMC10047013
- DOI: 10.3390/cells12060870
Knocking Down CDKN2A in 3D hiPSC-Derived Brown Adipose Progenitors Potentiates Differentiation, Oxidative Metabolism and Browning Process
Abstract
Human induced pluripotent stem cells (hiPSCs) have the potential to be differentiated into any cell type, making them a relevant tool for therapeutic purposes such as cell-based therapies. In particular, they show great promise for obesity treatment as they represent an unlimited source of brown/beige adipose progenitors (hiPSC-BAPs). However, the low brown/beige adipocyte differentiation potential in 2D cultures represents a strong limitation for clinical use. In adipose tissue, besides its cell cycle regulator functions, the cyclin-dependent kinase inhibitor 2A (CDKN2A) locus modulates the commitment of stem cells to the brown-like type fate, mature adipocyte energy metabolism and the browning of adipose tissue. Here, using a new method of hiPSC-BAPs 3D culture, via the formation of an organoid-like structure, we silenced CDKN2A expression during hiPSC-BAP adipogenic differentiation and observed that knocking down CDKN2A potentiates adipogenesis, oxidative metabolism and the browning process, resulting in brown-like adipocytes by promoting UCP1 expression and beiging markers. Our results suggest that modulating CDKN2A levels could be relevant for hiPSC-BAPs cell-based therapies.
Keywords: 3D culture; CDKN2A; UCP1; adipocytes; brown adipose progenitor; browning; human induced pluripotent stem cells; thermogenesis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

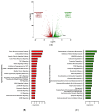
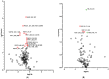

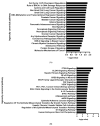
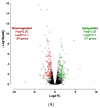



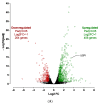

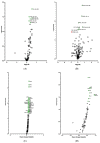

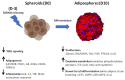
Similar articles
-
Cdkn2a deficiency promotes adipose tissue browning.Mol Metab. 2018 Feb;8:65-76. doi: 10.1016/j.molmet.2017.11.012. Epub 2017 Dec 1. Mol Metab. 2018. PMID: 29237539 Free PMC article.
-
Brown-like adipose progenitors derived from human induced pluripotent stem cells: Identification of critical pathways governing their adipogenic capacity.Sci Rep. 2016 Aug 31;6:32490. doi: 10.1038/srep32490. Sci Rep. 2016. PMID: 27577850 Free PMC article.
-
Brown-Like Adipocyte Progenitors Derived from Human iPS Cells: A New Tool for Anti-obesity Drug Discovery and Cell-Based Therapy?Handb Exp Pharmacol. 2019;251:97-105. doi: 10.1007/164_2018_115. Handb Exp Pharmacol. 2019. PMID: 29633179
-
White, brown, beige/brite: different adipose cells for different functions?Endocrinology. 2013 Sep;154(9):2992-3000. doi: 10.1210/en.2013-1403. Epub 2013 Jun 19. Endocrinology. 2013. PMID: 23782940 Review.
-
Brown and beige adipose tissue: a novel therapeutic strategy for obesity and type 2 diabetes mellitus.Adipocyte. 2021 Dec;10(1):48-65. doi: 10.1080/21623945.2020.1870060. Adipocyte. 2021. PMID: 33403891 Free PMC article. Review.
Cited by
-
The Role of β3-Adrenergic Receptors in Cold-Induced Beige Adipocyte Production in Pigs.Cells. 2024 Apr 19;13(8):709. doi: 10.3390/cells13080709. Cells. 2024. PMID: 38667324 Free PMC article.
-
Lactiplantibacillus plantarum N4 ameliorates lipid metabolism and gut microbiota structure in high fat diet-fed rats.Front Microbiol. 2024 Jun 7;15:1390293. doi: 10.3389/fmicb.2024.1390293. eCollection 2024. Front Microbiol. 2024. PMID: 38912346 Free PMC article.
-
PPARγ and C/EBPα enable adipocyte differentiation upon inhibition of histone methyltransferase PRC2 in malignant tumors.J Biol Chem. 2024 Oct;300(10):107765. doi: 10.1016/j.jbc.2024.107765. Epub 2024 Sep 12. J Biol Chem. 2024. PMID: 39276936 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

